White paper: Molecular healthcare
Genetic sequencing, next-generation sequencing, and the challenges and opportunities at hand

Molecular healthcare encompasses areas of industry development where we see significant opportunity over the next several years to improve healthcare outcomes and create substantial economic value.
While molecular healthcare includes genomics, lipidomics, proteomics, transcriptomics, microbiomics, and metabalomics, to name a few, in this report we focus on the fast-moving area of genomics and the early-innings impact next-generation sequencing (NGS) is having on the proliferation of molecular diagnostics in the clinical setting.
We estimate the NGS platform technology market to be ~$2.3 billion today and growing 20% annually with a total addressable market of ~$45 billion. This report reviews the market size, technology platforms, emerging trends, regulatory and reimbursement issues, as well as key growth drivers and moderators for NGS and the ~$7.7 billion molecular diagnostic industry.
We also review genetic sequencing basics and profile the key publicly traded and privately held innovators and major players.
TABLE OF CONTENTS
Defining molecular healthcare
The term “molecular healthcare” is one we use to describe the current impact and direction we see genomics and ancillary molecular testing taking in medicine. While other terms like “precision” or “personalized medicine” are used to describe the advances genomics and other technologies have made in healthcare, we contend the goals of precision or personalized medicine have yet to be achieved. We are of the view that the current practice of medicine remains an inexact science and still tends to be reactive rather than preventive, predictive, and accurate. Therefore, we feel “molecular healthcare” better characterizes the current state of genomics in medicine.
While we feel we are still in the early innings of unlocking the true potential of genomics, recent discoveries from sequencing the human genome and movement into the clinical setting have enhanced researchers’ ability to classify individuals into subpopulations by disease type or treatment method. In our view, the further development and use of molecular healthcare will be fueled by healthcare consumers gradually gaining a better understanding of the meaning and value of molecular information as well as clinicians being armed with the right tools and technologies to deal with the complexity of disease diagnosis, onset, progression, treatment, prognosis, and outcome.
DNA structure
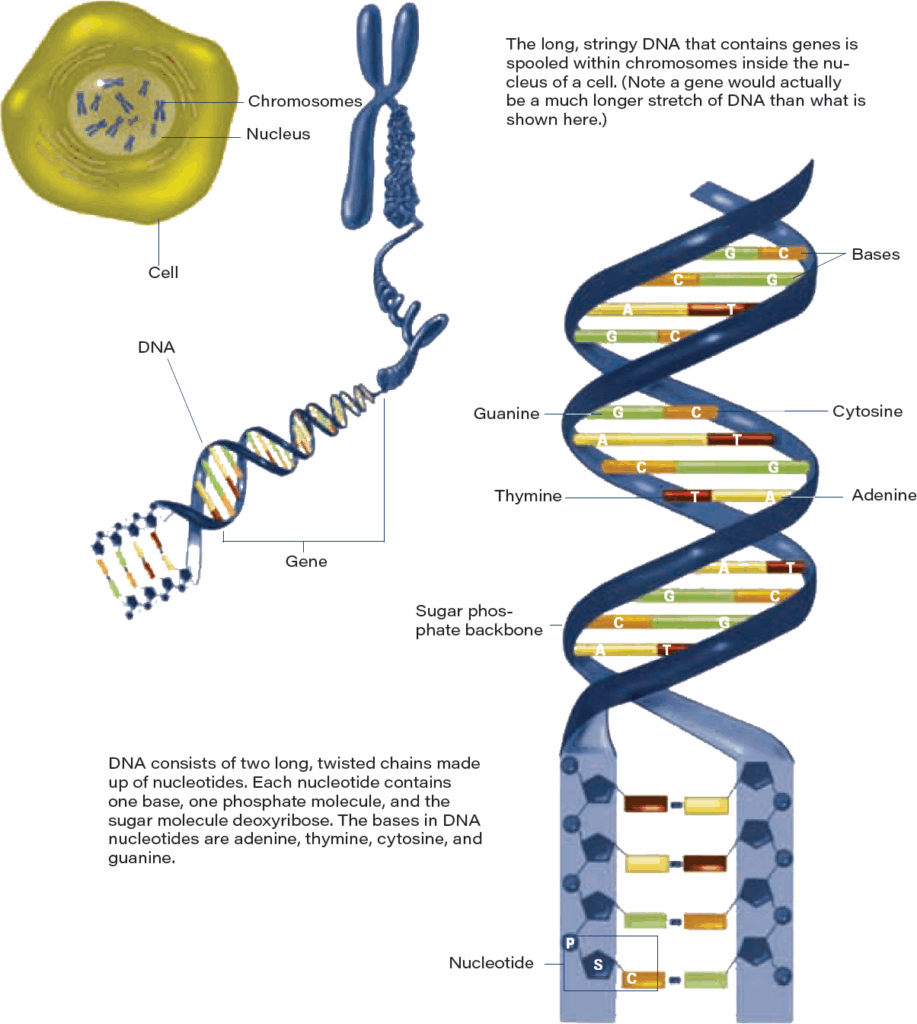
Source: U.S. Department of Health and Human Services, The New Genetics.
To achieve the ultimate goal of precision or personalized medicine, further technological advances and infrastructure changes must be realized in the areas of sequencing, testing, and regulation. Although we think we are only in the first couple innings of the development of genomics applications in healthcare, the foundation for such an ecosystem is already being formed, and we believe molecular healthcare is a key early pillar leading to personalized healthcare.
Fundamentals of genetic sequencing
Molecular OS
Much like Google’s Android or Apple’s iOS, the human genome is an operating system with numerous application possibilities and, on the surface, seems relatively simple to analyze and interpret. However, extracting, reassembling, and understanding this code is extremely complex and a major challenge. By definition, a genome is all the genetic material of an organism. It consists of deoxyribonucleic acid (DNA), which contains the biological instructions. DNA, along with the instructions it contains, is passed from adult organisms to their offspring during reproduction.
DNA is made of chemical building blocks called nucleotides. These building blocks are made of three parts: a phosphate group, a sugar group, and one of four types of nitrogen bases. To form a strand of DNA, nucleotides are linked into chains, with the phosphate and sugar groups alternating. The four types of nitrogen bases found in nucleotides are adenine (A), thymine (T), guanine (G), and cytosine (C). The sequence of these bases determines the information available for building and maintaining an organism, similar to the way in which letters of the alphabet appear in a certain order to form words.
Difference between DNA and RNA
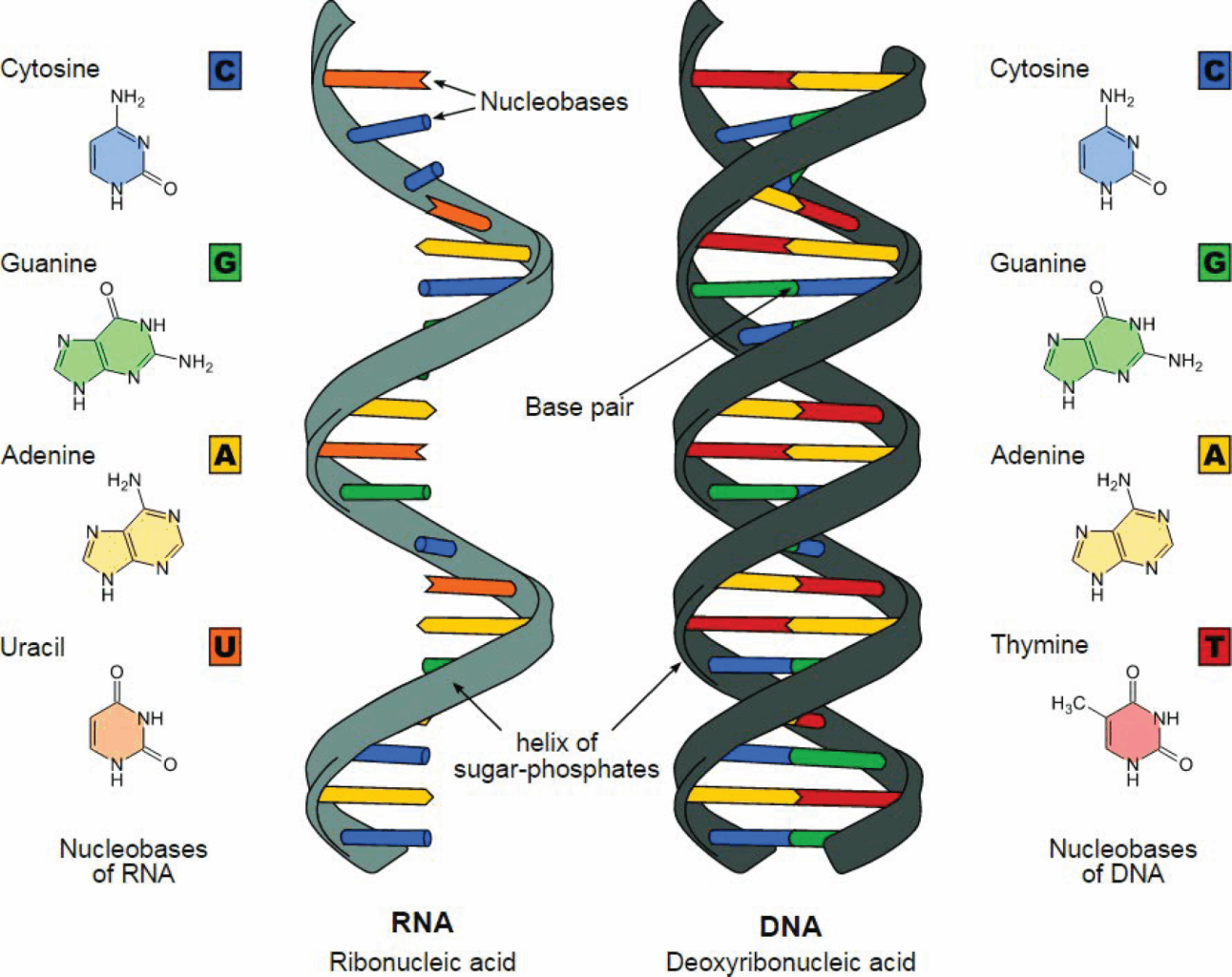
Source: Wikipedia.
For example, the sequence ATCGTT might instruct for blue eyes, while ATCGCT might instruct for brown. The complete DNA genome of a human contains about three billion base pairs of only four repeating bases. Of those bases, 99% are the same in all people. These three billion base pairs are divided into 23 chromosomes ranging in size from 50 million to 250 million bases. Contained within these chromosomes are approximately 23,000 smaller regions, called genes, each one containing the recipe for a protein or a group of related proteins.
Differences between genomes at different positions can be highly interactive; for example, a mutation that increases the lifetime risk of cancer in one genomic context may decrease risk in another context. While the two copies of the 23 chromosomes we as humans inherit are important in determining who we are, our genomes continue to change as we age. Understanding the genetic makeup of an individual, including localized mutations that take place after conception, is a key to understanding and treating diseases.
Importance of DNA sequencing
When a gene is expressed, a partial copy of its DNA sequence, called RNA, is used as a template to direct the synthesis of a protein, which in turn directs all cellular functions. Ribonucleic acid (RNA) is a molecule similar to DNA. However, unlike DNA, RNA is single-stranded. An RNA strand has a backbone made of alternating sugar (ribose) and phosphate groups. Attached to each sugar is one of four bases–adenine (A), uracil (U), cytosine (C), or guanine (G). Different types of RNA exist in the cell, including messenger RNA (mRNA), ribosomal RNA (rRNA), and transfer RNA (tRNA).
Researchers use sequencing to determine the structure and order of a given DNA fragment. This sequence of DNA encodes the necessary information for that living organism to survive and reproduce. Variations among organisms are due in large part to differences in their DNA sequences. Variations in the sequences due to insertions, deletions, or duplications of nucleotide bases may result in certain genes becoming overexpressed, underexpressed, or silenced altogether, which could trigger changes in cellular function. Determining the sequence of DNA helps researchers determine why and how organisms function. Once scientists have determined the DNA sequence of an organism, they use this knowledge to identify, diagnose, and potentially develop treatments for genetic diseases.
Defying Moore’s law
The Human Genome Project (HGP) took $1 billion and 13 years to sequence the first draft of the human genome. At the conclusion of the HGP, the National Human Genome Research Institute of the National Institutes of Health published a vision for the future of genomics research. This vision included technology development as a centerpiece and articulated the goal of sequencing an entire genome for $1,000. In addition, clinical use of genomics needed to be highly accurate, rapid, and far less expensive. In the 12 years since the completion of the HGP, advances in genome technology have led to an exponential decrease in sequencing costs, declining at a rate that exceeds Moore’s law.
TABLE 1: Sequencing cost per genome
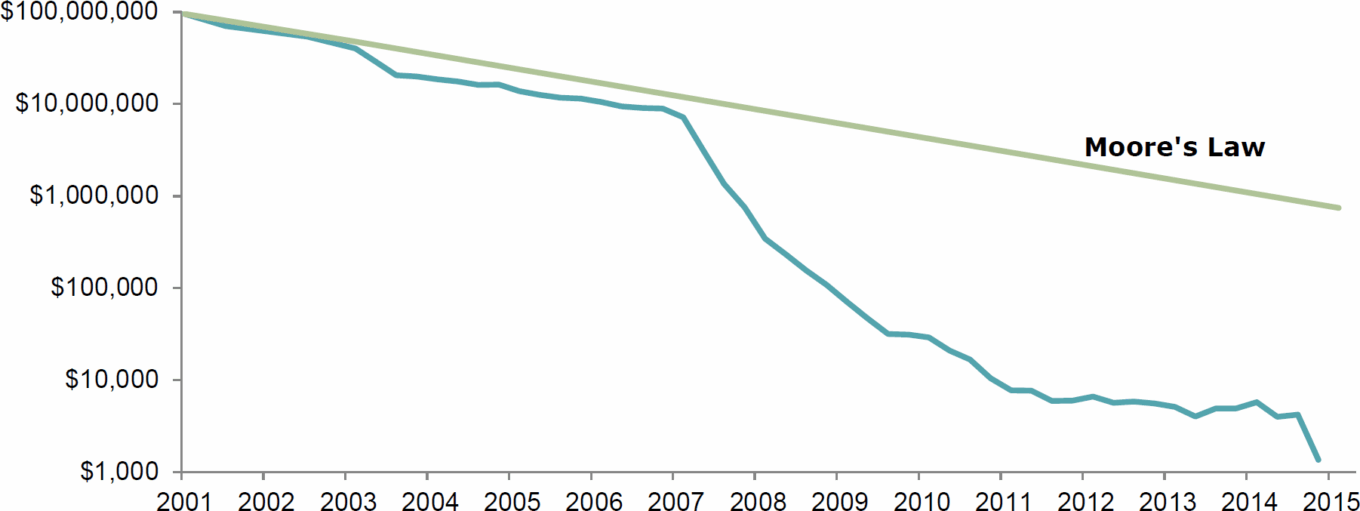
Source: National Institute of Health.
In January 2014, Illumina turned the goal of the $1,000 genome into reality, or nearly so, with the launch of its HiSeq X Ten system.
Reaching the $1,000 benchmark was a major breakthrough, in our view, because it is comparable to costs of existing medical tests and procedures and has begun to attract a market for clinical tests based on next-generation sequencing (NGS). When fully deployed in the clinic, the tools of DNA sequencing have the promise of predicting disease susceptibility, disease course, optimal treatment strategies, and disease prevention.
TABLE 2: Calculating the $1,000 human genome (January 2024)
| HiSeq X Ten | |
|---|---|
| Consumable reagent cost per genome | $797 |
| Hardware amortization rate per genome | $137 |
| DNA extraction, sample prep & labor | $66 |
| Total cost per human genome sequenced | $1,000 |
Source: First Analysis, Illumina.
Variations in the human genome
Many different types of DNA variation can occur among human genomes, which, together with environmental factors, account for differences among individuals. These variations can range from single nucleotide changes to gain or loss of whole chromosomes, and they may be inherited or occur spontaneously (de novo). Many DNA variants arise from a failure to repair damage or correct replication errors and as a result of recombination events.
The most common form of variation in humans is called a single-nucleotide polymorphism (SNP), also known as a point mutation or single-base substitution. SNPs occur every 1,000 bp (base pairs) on average throughout the genome. SNPs that occur in more than 1% of the population are classified as common variants; these are often located in noncoding regions and tend to have little or no phenotypic effects. SNPs that occur in less than 1% of the population are classified as rare variants or mutations, and they may be associated with a pharmacological interaction or to the risk of developing certain diseases.
Single nucleotide polymorphism
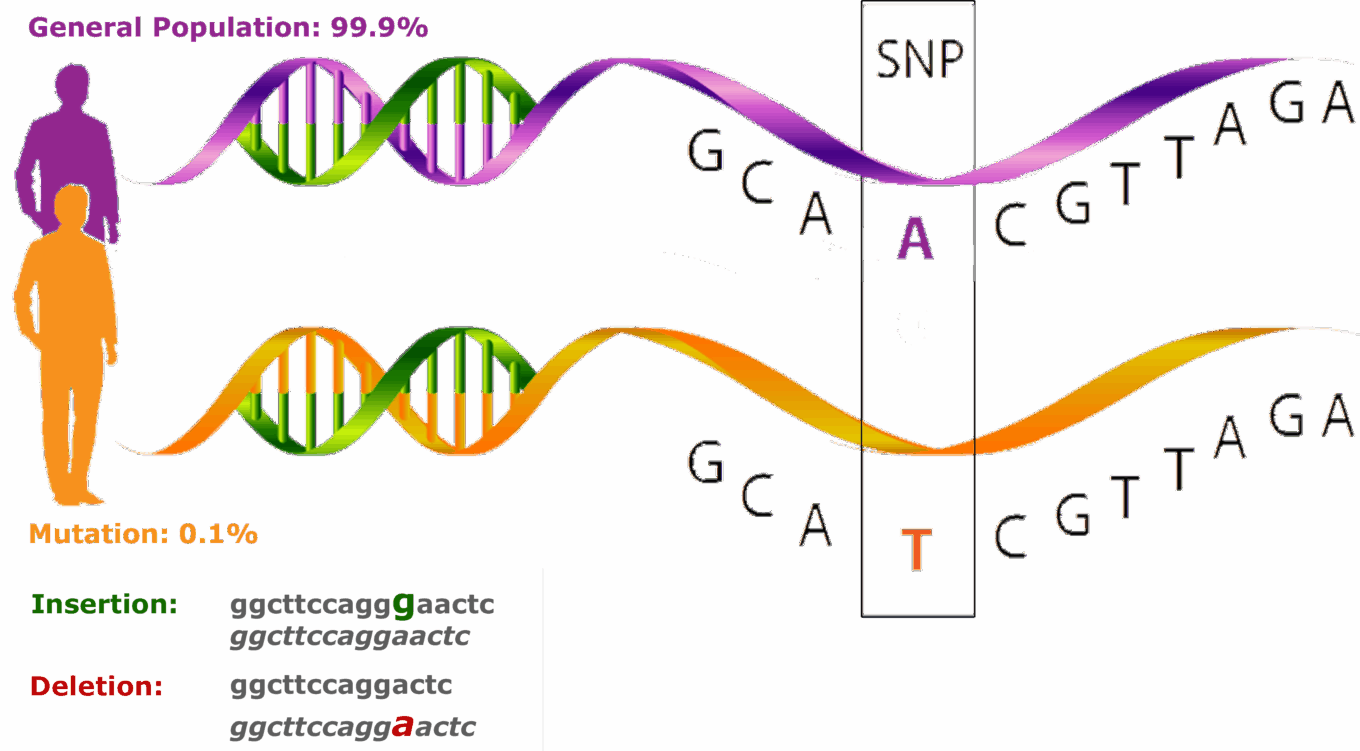
Source: neuroendoimmune.files.wordpress.com.
Genomic variation can also be caused by multiple base changes, the most common of which is in the form of INDELs—INsertions and DELetions that range in size from 1 to 1,000 base pairs. Variations in the number of a gene are referred to as copy number variants (CNVs) and include both common and rare variants.
When large areas of a gene or chromosome (or entire chromosomes) are deleted or duplicated, this can lead to changes in the level of expression of genes in the area affected, and the changes are important in the development and progression of diseases.
In addition to variation that occurs at the DNA sequence level, epigenetic variation also contributes to both nonheritable and heritable differences in gene expression. Epigenetics is the study of changes in the phenotype that are caused by external or environmental factors that switch genes on and off and affect how cells read genes instead of being caused by changes in the DNA sequence. Examples of epigenetic variations include DNA methylation and histone modifications, each of which alters how genes are expressed without altering the underlying DNA sequence.
Epigenetic mechanism
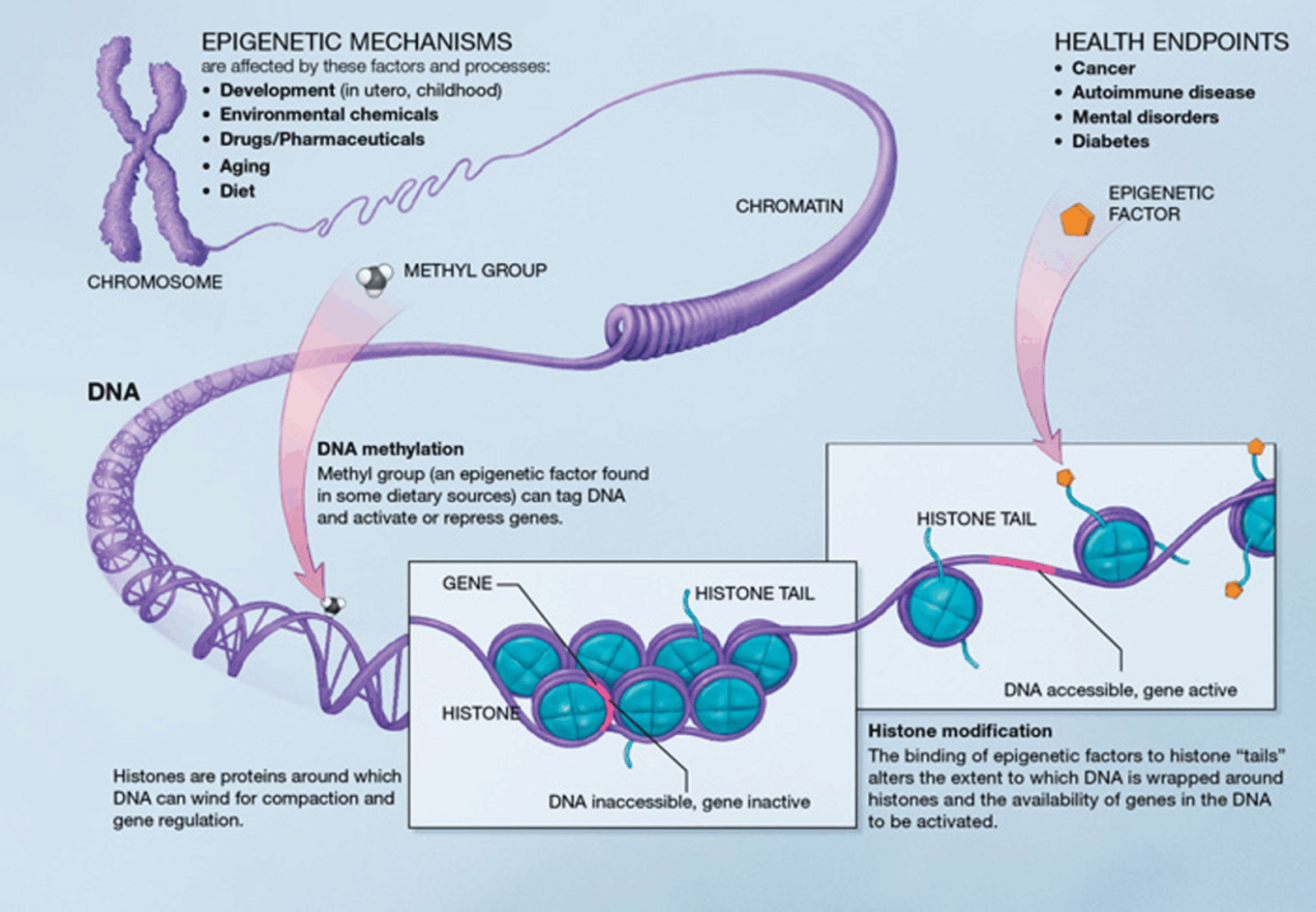
Source: National Institute of Health.
Sanger sequencing, developed in the 1970s by Frederick Sanger, was the most widely used method of DNA sequencing and was used to sequence the first genome in 1977 as well as in the HGP. During sample preparation for Sanger sequencing, scientists construct different-sized fragments of DNA, each starting from the same location. Each fragment ends with a particular base that is labeled with one of four fluorescent dyes corresponding to that particular base. Then, all of the fragments are organized in order of their length by driving them through a gel. Information regarding the last base is used to determine the original sequence. Under standard conditions, this method results in a read length that is approximately 700 bases on average but may extend to 1,000 bases. These are relatively long read lengths compared with many other sequencing methods.
However, Sanger sequencing is limited by the small amounts of data that can be processed per unit of time, referred to as throughput. Sanger sequencing has since been supplanted by “next-generation sequencing” in many applications, especially for large-scale, automated genome analyses. However, the Sanger method remains in wide use for smaller-scale projects, for validation of NGS results, and for obtaining especially long contiguous DNA sequence reads (>500 nucleotides) mainly due to its superior accuracy.
Evolution of DNA sequencing technologies
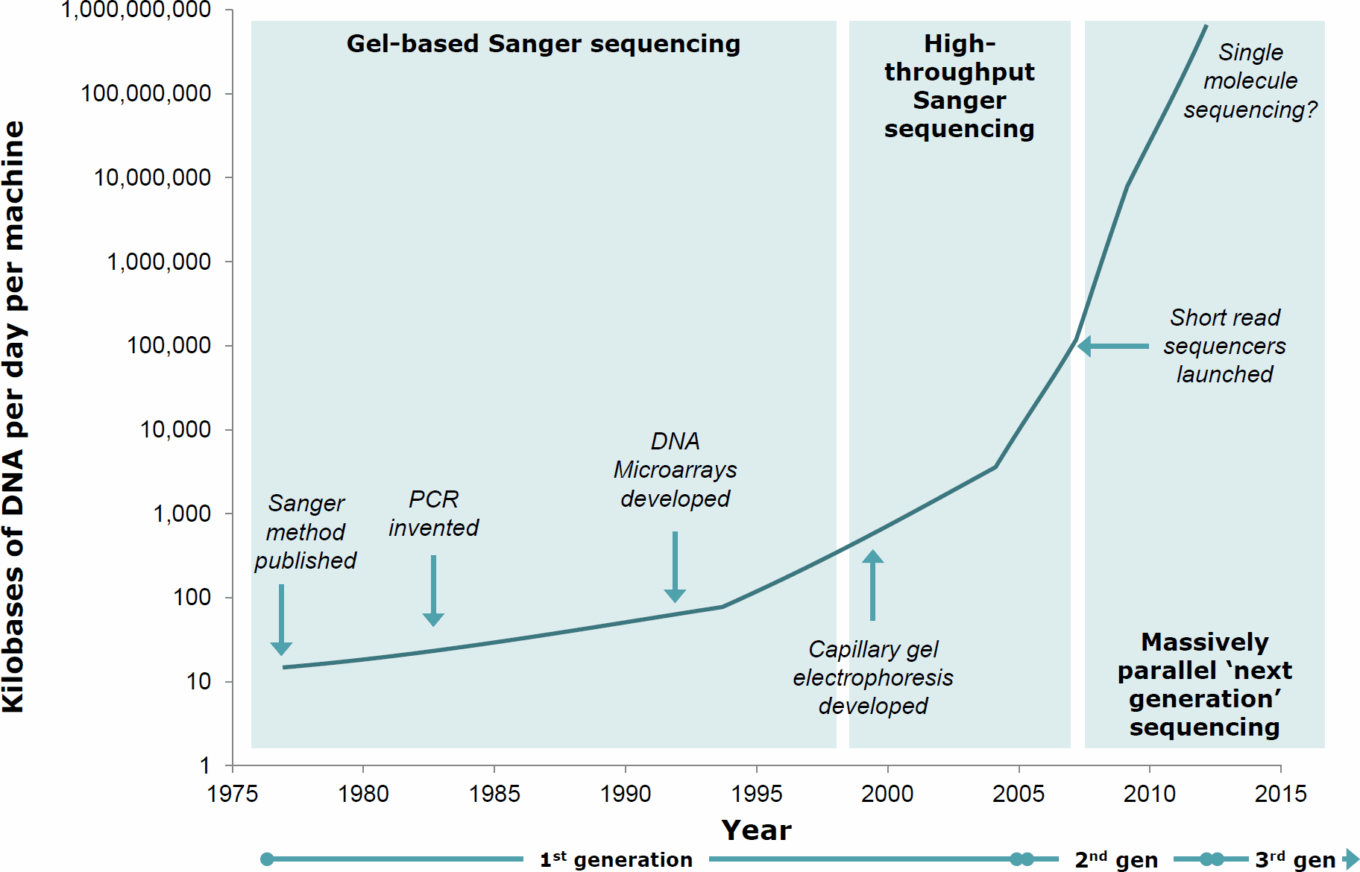
Source: PHG Foundation, Next Steps in the Sequence..
Next-generation sequencing, or high-throughput sequencing, was introduced commercially in 2005 in response to the low throughput limitations of Sanger sequencing. The main difference between Sanger sequencing and NGS is that NGS sequences millions of small DNA fragments at the same time or in parallel and therefore dramatically increases the throughput per reaction, making it more suitable for addressing the challenges associated with the increasing demands for testing of multiple gene markers. This key innovation enables screening of large numbers of genes or samples while keeping short turnaround time for timely clinical reporting. More importantly, screening multiple markers with NGS technology requires a single input of relatively low-quantity DNA or RNA in contrast to Sanger sequencing, which needs cumulatively larger quantities of genetic material. NGS also decreases the overall cost of multiple-marker screening compared with the costs of Sanger sequencing.
Sanger sequencing
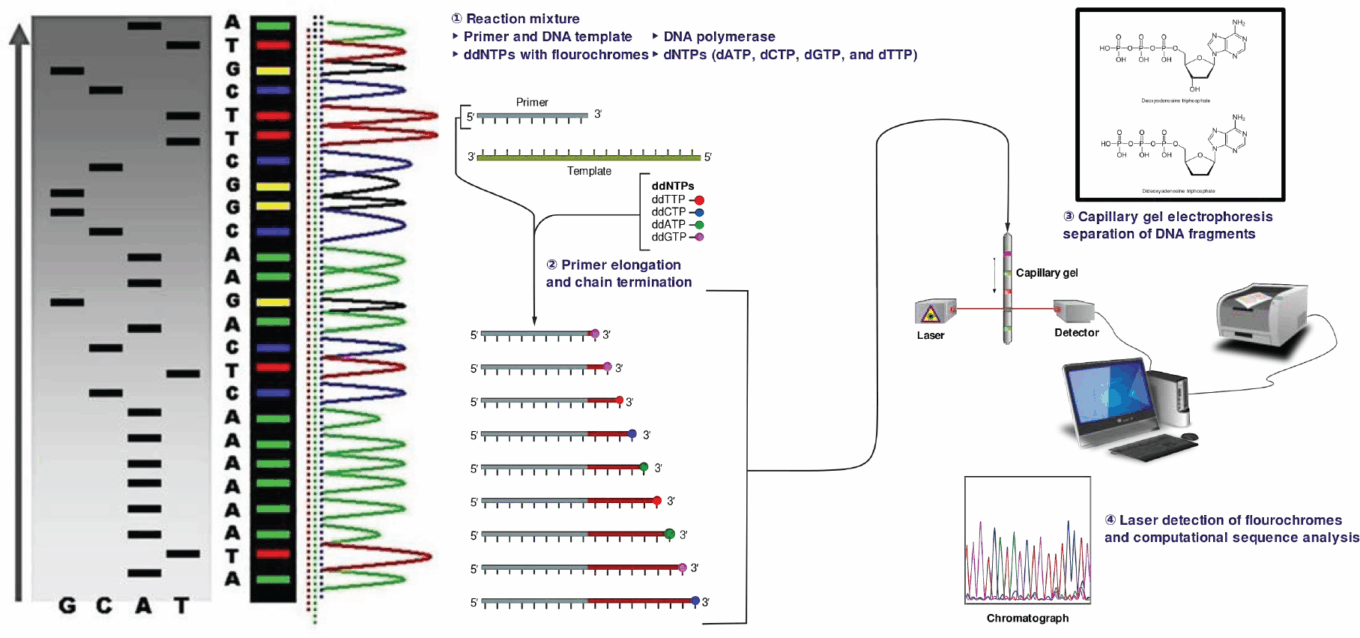
Source: Wikipedia.
There are three fundamental steps in all NGS sequencing formats: preparation, immobilization, and sequencing. Sample preparation involves random fragmentation of DNA and the addition of adapter sequences to the ends of the fragments. The prepared fragments are then immobilized on a solid support to form detectable sequencing features. Finally, massively parallel cyclic sequencing reactions are performed to decipher the nucleotide sequence.
High-throughput sequencing workflow
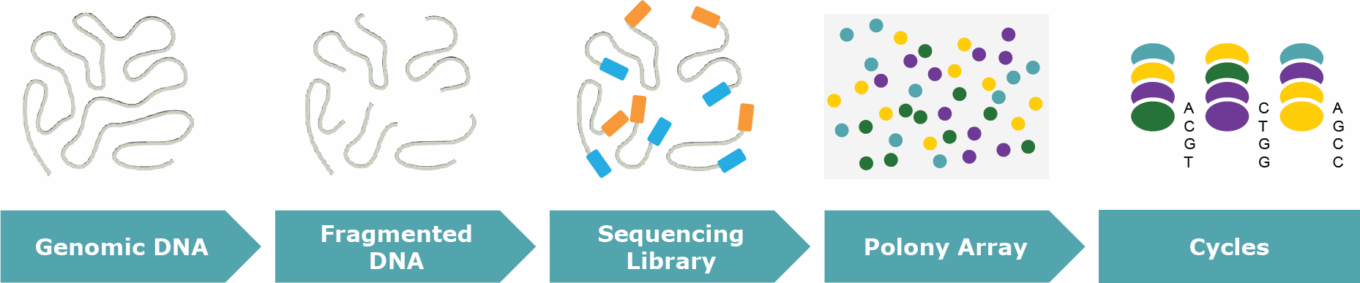
Source: First Analysis.
Next-generation sequencing
Common NGS applications
Whole-genome sequencing
Whole-genome sequencing (WGS) involves sequencing all of the base pairs in the genome. Although it seems logical that if the “whole genome” is sequenced, then all variants will be detected, this is not technically true. Some variants can be missed due to variation in coverage across the genome, and in some cases these “missed” variants can be detected by whole-exome sequencing (WES) due to the higher coverage achieved in certain areas with target-enriched sequencing. Therefore, as neither WES nor WGS detects all variants, some researchers advocate performing both exome and genome sequencing on the samples to ensure that as many variants as possible are detected. The main advantage of WGS is it can detect structural aberrations that occur outside exonic areas in the genome, such as translocations and rearrangements. Additionally, variations occurring in DNA regions containing regulatory elements, such as enhancers or silencers, can only be analyzed by WGS.
Exome sequencing
Exome sequencing, or WES, is a targeted sequencing technique for all the protein-coding genes in a genome (known as the exome). It consists of first selecting only the subset of DNA that encodes proteins (known as exons) and then sequencing that DNA using any NGS technology. There are 180,000 exons, which constitute about 2% of the human genome, or approximately 30 million base pairs, but mutations in these sequences are felt to be much more likely to have severe consequences than in the remaining 98%. The goal of this approach is to identify genetic variation that is responsible for both Mendelian and common diseases such as Miller syndrome and Alzheimer’s disease without the high costs associated with whole-genome sequencing. Although more cost-effective than WGS, WES is not able to identify structural and noncoding variants associated with disease, which can be found using WGS. WES provides less data than WGS, but WES datasets are easier to interpret because the clinical consequences of alterations in intronic regions captured by WGS are not yet understood in terms of phenotype and disease.
Targeted sequencing
In targeted sequencing, a subset of genes or regions of the genome are isolated and sequenced. Targeted sequencing allows researchers to focus on specific geographies of DNA that are most relevant and enables sequencing at higher coverage levels. Compared to WGS, targeted sequencing is a more cost-effective method for investigating areas of particular interest.
TABLE 3: Target enrichment systems for NGS
| Company | Enrichment technology | Approach |
|---|---|---|
| Illumina | TruSeq | DNA probe-based capture |
| Life Technologies (Thermo Fisher) | AmpliSeq | PCR-based amplifications |
| Agilent Technologies | SureSelect | Hybridization and capture using cRNA-baits |
| Agilent Technologies | Haloplex | Circularization probe-based target enrichment |
| Qiagen | GeneRead | PCR-based amplification |
| RainDance Technologies | ThunderStorm and ThunderBolt systems | DNA probe-based capture |
| Integrated DNA Technologies | Xgen Lockdown probes | Droplet PCR_based amplification |
Source: First Analysis, company reports.
De novo sequencing
In de novo sequencing, there is no pre-existing sequence to search against. So the reads must be searched against each other to determine overlaps and join them into contiguous segments of sequence (called “contigs”). The alignment will be conducted progressively, with each read that overlaps another being replaced by a contig sequence. Like a jigsaw puzzle, the longer the contig, the easier it is to assemble the sequence.
TABLE 5A: NIH budget ($ in millions)
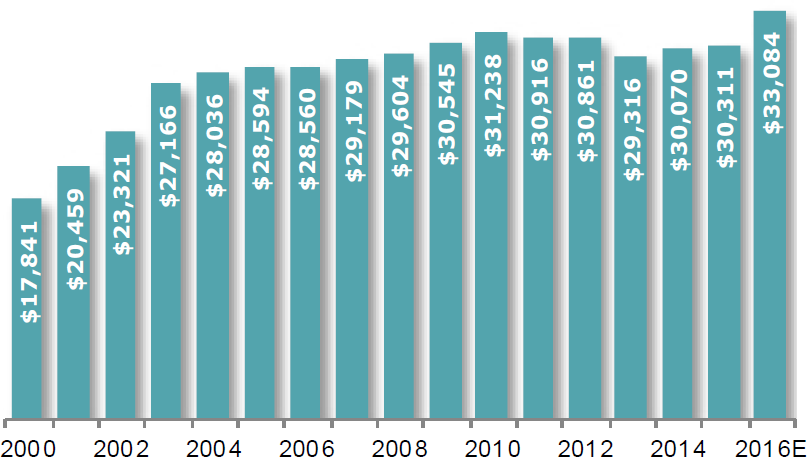
TABLE 5B: Global pharma R&D spending ($ in millions)
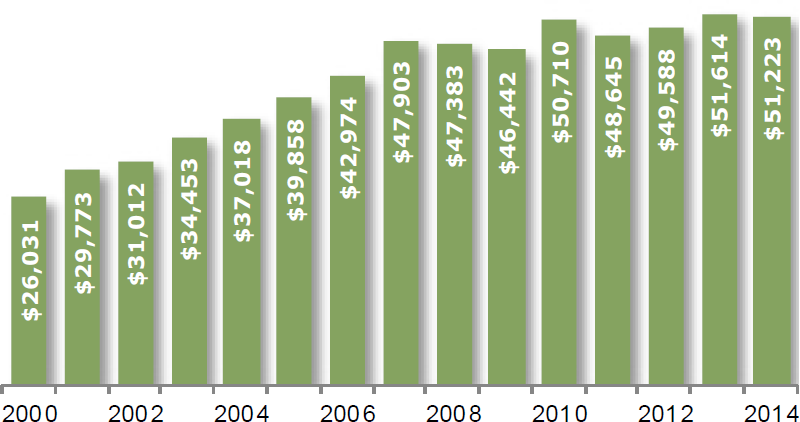
Source: First Analysis, National Institute of Health, PhRMA.org.
NGS market dynamics
Over the years, the sequencing industry has gone through periods of excitement and/or hype, followed by periods of disappointment as advances have not materialized as quickly as hoped or promised. The sheer potential of NGS has attracted many companies, researchers, investors, and others, which has resulted in NGS becoming a large market in a relatively short time. Valued today at roughly $2.3 billion, the NGS market by our estimate will grow at a 20% CAGR through 2022. We see the growth being driven by continuous technological innovations aimed at higher throughput, increased accuracy, lower costs, growing clinical adoption, and drug development. In addition, we see President Obama’s Precision Medicine Initiative, coupled with an expected $2-$3 billion increase in NIH funding slated for 2016 (on a base of ~$30 billion), further benefiting the industry. We estimate ~35% of NGS total market revenue is derived from the academic/research setting.
TABLE 5: 2015 NGS data
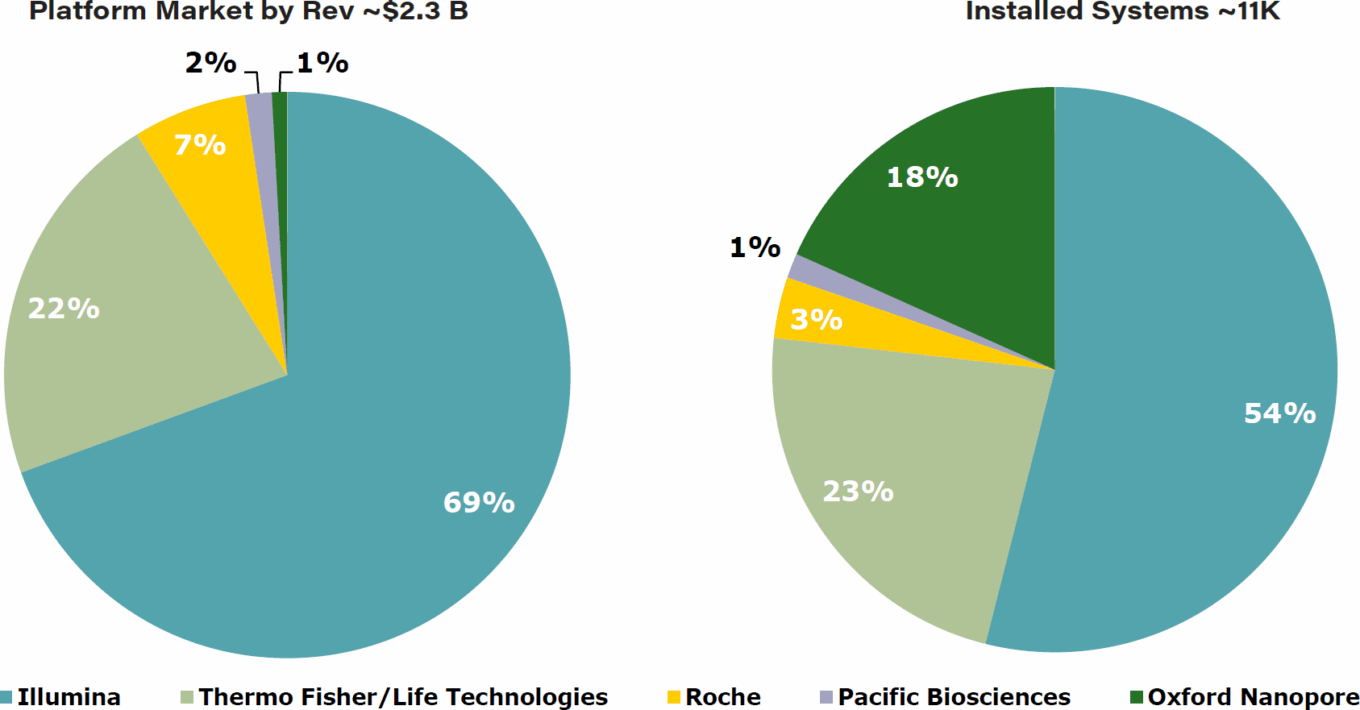
Source: First Analysis, company reports.
Currently, the NGS market leader is Illumina by both revenue and systems in the marketplace, and while the sequencing market is constantly evolving and increasingly competitive, we think Illumina, with its robust product platform and large installed base, is well positioned to maintain its leadership and bellwether share of the market.
TABLE 6: NGS total addressable market ~$45 billion
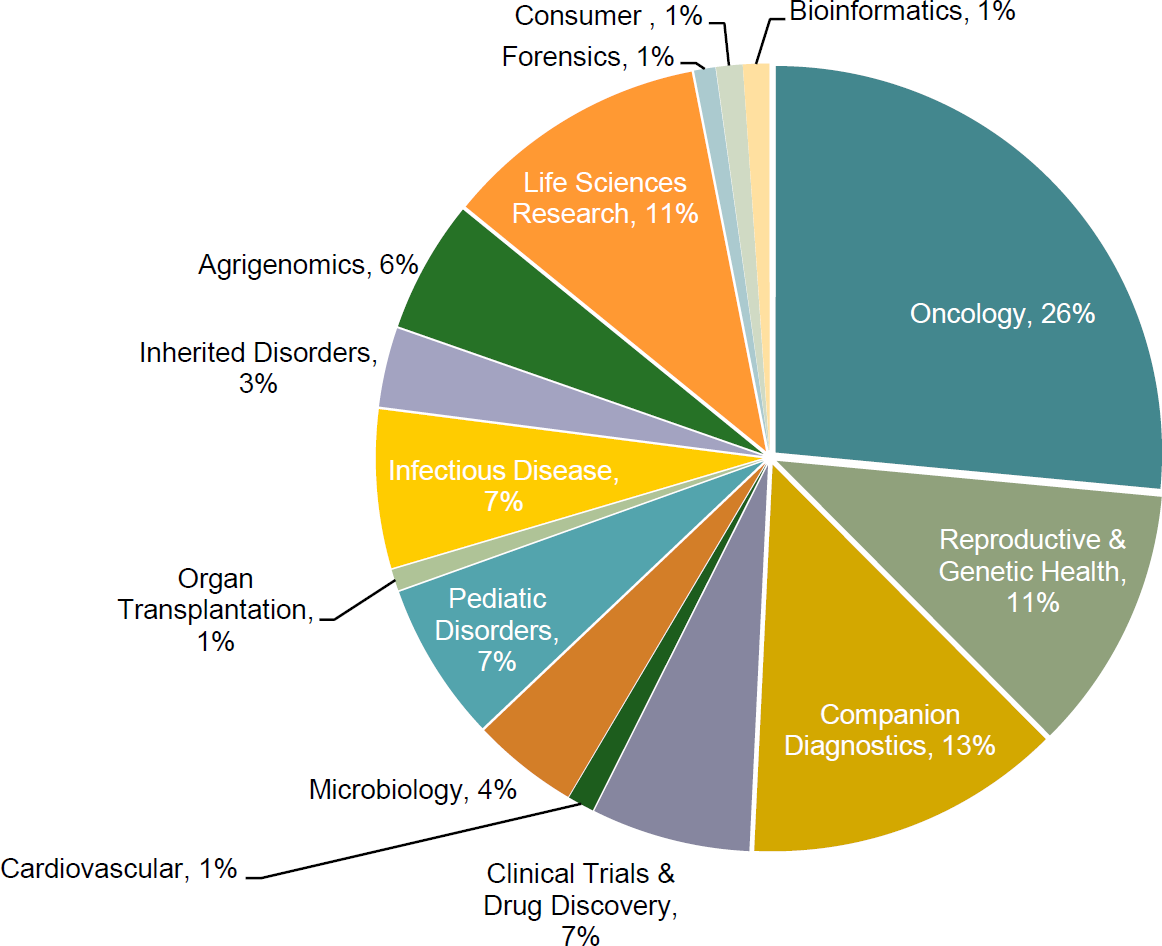
Source: First Analysis, Bloomberg, company reports.
Clinical adoption of NGS
Despite the massive potential opportunity NGS presents and the falling instrumentation costs, expansion and adoption outside the research/academic setting and into other end-user markets, specifically the clinical setting, have been slower than many in the industry had anticipated. In our view, core moderators to broad clinical adoption are ambiguous findings reported by the research community are not clinically actionable or conclusive, and there is no uniform therapeutic or payer reimbursement available. In order to have widespread adoption of NGS into the clinical practice, it must be rigorously tested for regulatory approval, accepted and understood by both clinicians and payers as a routine diagnostic tool, and implemented into clinical trials to ultimately improve patient outcomes and medical economics.
Despite the challenges, we expect clinical adoption of NGS to continue primarily in the form of diagnostic tests incorporated into routine practice. One such example is Illumina’s MiSeqDx instrument, which became the first NGS instrument approved by the FDA for use in clinical diagnostics. While we don’t expect rapid clinical adoption to occur, we see declining sequencing technology costs, reimbursement progress, expanded test offerings, faster turnaround times, and lower overall treatment costs as catalysts to further clinical utilization of NGS over the next five years or so.
NGS technology platforms
All NGS platforms share a common technological feature—massively parallel sequencing of clonally amplified or single DNA molecules that are spatially separated in a flow cell. These platforms are briefly summarized below.
454 (Roche)
454 Life Sciences was as founded in 2000 originally as 454 Corp., a subsidiary of CuraGen Corp., and was acquired by Roche in 2007 for ~$155 million. 454 brought the first next-generation sequencing technologies to market in 2005. In October 2013, Roche announced it would shut down 454 and stop supporting the platform by mid-2016. The overall sequencing approach for 454 systems is pyrosequencing, a DNA sequencing method based on the “sequencing by synthesis” (SBS) principle. The pyrosequencing SBS method is based on detecting the activity of DNA polymerase (a DNA synthesizing enzyme) with another chemiluminescent enzyme. Essentially, the method allows sequencing of a single strand of DNA by synthesizing the complementary strand along it, one base pair at a time, and detecting which base was actually added at each step. The template DNA is immobile, and solutions of A, C, G, and T nucleotides are sequentially added and removed from the reaction. Light is produced only when the nucleotide solution complements the first unpaired base of the template. The sequence of solutions that produce chemiluminescent signals allows the determination of the sequence of the template. One shortcoming of the 454 approach is it frequently misidentifies the length of homopolymers (stretches of repeats). Additionally, this technology is often considered not cost-effective when compared with other next-generation sequencing technologies.
In an effort to strengthen its NGS pipeline, Roche, in 2014, made a strategic investment of ~$15 million in nanopore sequencing startup Stratos Genomics, which uses a sequencing method called sequencing by expansion (SBX). SBX converts DNA into more easily read polymer and, according to the company, makes it cheaper and faster to sequence DNA. In addition, Roche acquired Genia Technologies, a nanopore sequencing platform, for $125 million in cash and up to $225 million in milestone payments. At the heart of Genia’s technology is a single-molecule semiconductor DNA sequencing platform using nanopore technology.
Illumina (Solexa / Lynx Therapeutics)
At the heart of Illumina’s industry-leading sequencing technology is its SBS technology. Two Cambridge University scientists, Shankar Balasubramanian and David Klenerman, first developed Illumina’s version of SBS technology in the mid-1990s. They used fluorescently labeled nucleotides to observe the motion of polymerase as it synthesized DNA immobilized to a surface.
The steps of sequencing by synthesis (SBS)
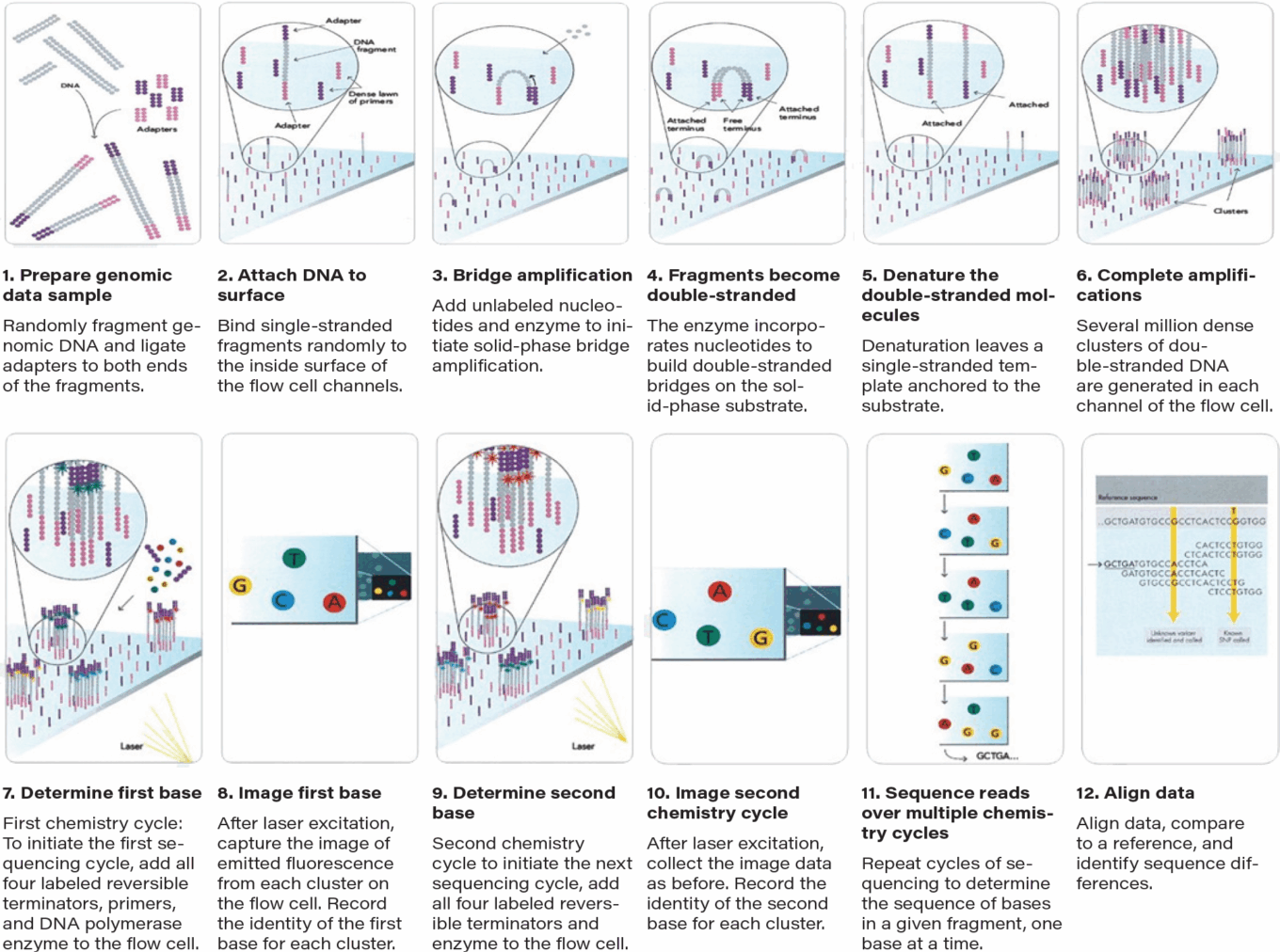
Source: Illumina.
In 1998, Balasubramanian and Klenerman obtained venture capital funding and formed Solexa. For the next eight years, Solexa developed and acquired new molecular technologies to enhance its molecular sequencing platform. In 2006, Solexa launched its first sequencer, the Genome Analyzer, which gave scientists the power to sequence one billion bases in a single run.
In early 2007, Illumina acquired Solexa for its SBS technology. SBS is the most successful and widely adopted next-generation sequencing platform worldwide. SBS technology uses four fluorescently labeled nucleotides to sequence the tens of millions of clusters on the surface of Illumina’s proprietary flow cell. During each sequencing cycle, a single labeled deoxynucleoside triphosphate (dNTP) is added to the nucleic acid chain. Once the dNTP is combined, the fluorescent dye is imaged to identify the base and then cleaved to allow the integration of the next nucleotide. This process is run hundreds of millions of times in order to deliver high sequencing output and a fast data generation rate.
TABLE 7: Illumina sequencing solutions
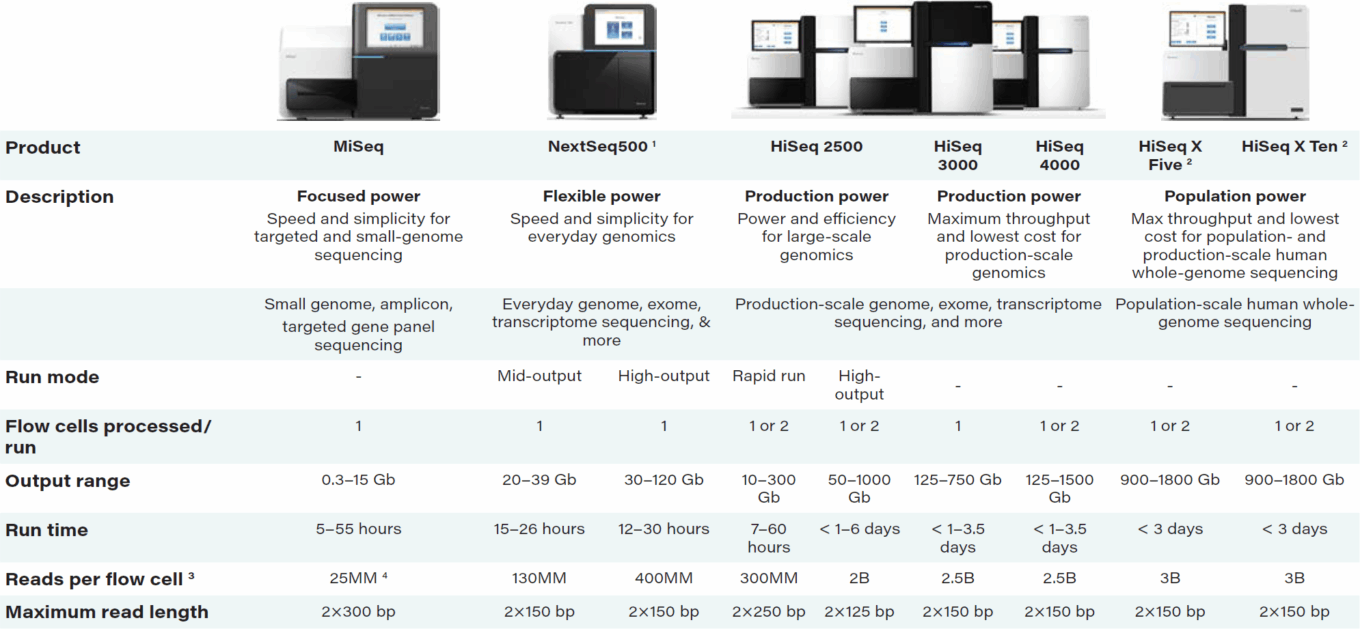
Source: Illumina.
Notes: (1)The NextSeq 550 System has identical sequencing specifications to the NextSeq 500 System and includes array scanning functionality for cytogenomic and karyomapping applications. (2) Specifications shown for an individual HiSeq X System. The HiSeq X System is available only as part of the HiSeq X Five or HiSeq X Ten System. (3) Clusters passing filter. (4) For MiSeq Reagent Kits v3 only.
TABLE 8: Illumina sequencing cost metrics
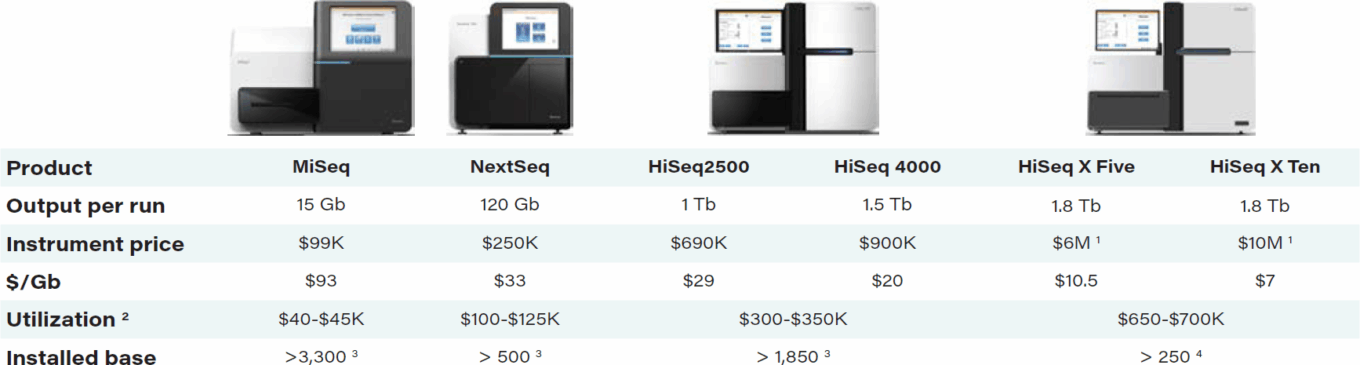
Source: Illumina.
Notes: (1) Price based on system of 5 and 10 units for X Five and X Ten, respectively. (2) Company’s projected annual instrument utilization. (3) Based on end of fiscal 2014. (4) Based on end of Q3 2015.
Thermo Fisher Scientific
(Life Technologies / Ion Torrent / Applied Biosystems)
Life Technologies, acquired by Thermo Fisher in 2014 for ~$16 billion, itself was the result of the $6 billion merger of Invitrogen and Applied Biosystems (SOLiD) in 2008. In 2010, Life Technologies acquired Personal Genome Machine (PGM) maker Ion Torrent for $750 million. After the 2014 acquisition, Thermo Fisher integrated Life Technologies into its Life Sciences Solutions segment. Currently, Thermo has two NGS platforms, SOLiD and Ion Torrent.
SOLiD sequencing involves a ‘’sequencing-by-ligation’’ chemistry approach as opposed to the ‘’sequencing-by-synthesis’’ method used by Illumina. The ligation method of sequencing provides internal accuracy checks as each ligation is coded by two nucleotides. As a result, each nucleotide is sequenced twice, and the overall accuracy of the sequencing data is 99.9%—one of the highest on the market. Despite its high accuracy rate, then-Life Technologies had trouble placing SOLiD instruments due to customer complaints the workflow was too complicated. While Thermo still sells the SOLiD sequencing platform, currently offering two high-throughput models, the 5500 and the 5500xl, the company has turned its NGS focus toward Ion Torrent.
TABLE 9: SOLiD sequencers
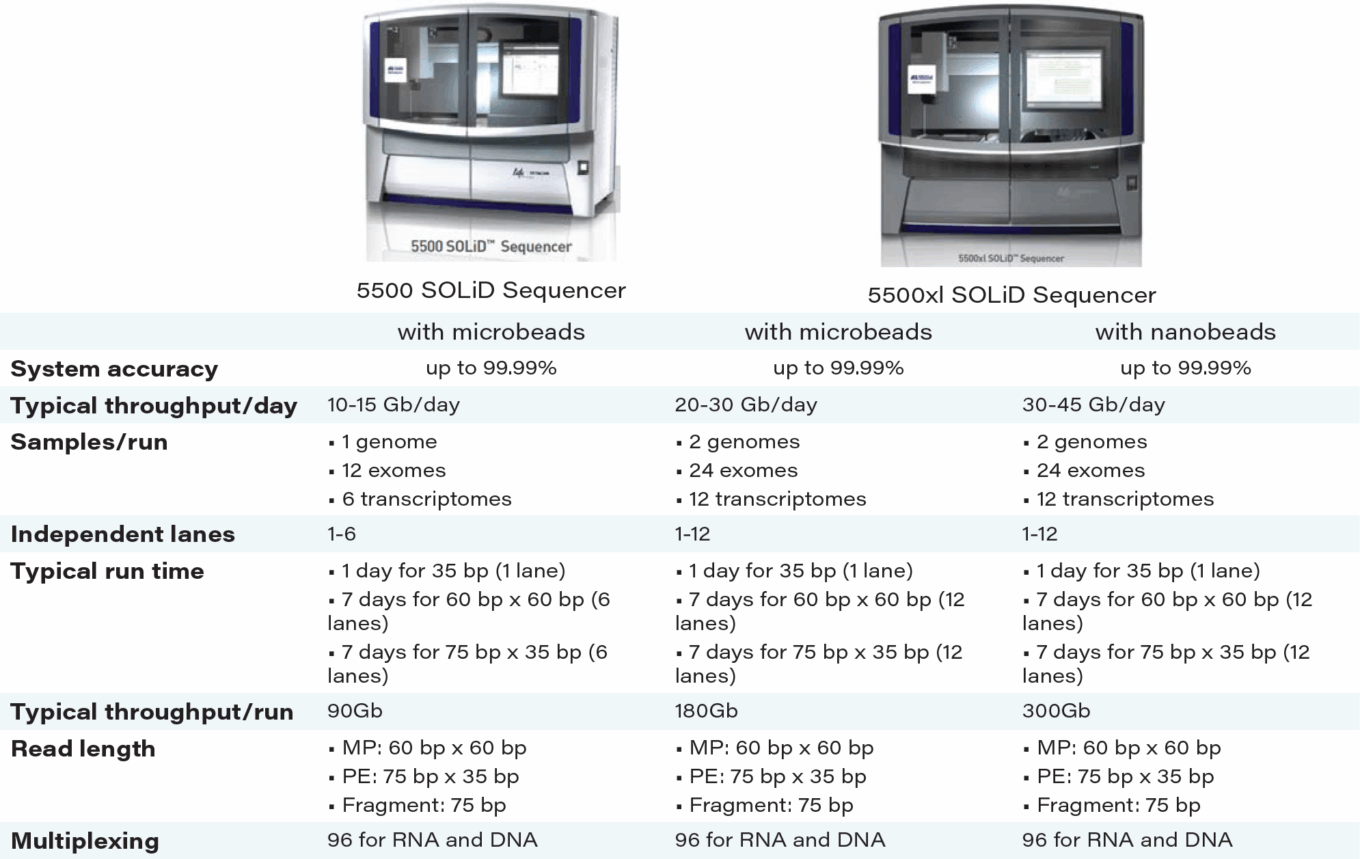
Source: Thermo Fisher Scientific.
Ion Torrent entered the NGS market in 2010 with the Personal Genome Machine (PGM). The Ion Torrent sequencing platform is based on the sequencing-by-synthesis method and uses proprietary semiconductor technology to monitor the synthesis reaction. Unlike other platforms, sequencing is based on monitoring the release of hydrogen protons, which are by-products of DNA synthesis. In a nutshell, DNA fragments are held in specialized semiconductor-chip-based microwells containing beads to which the fragments are attached. The microwells are designed to be sensors, and nucleotides are added sequentially to each microwell. If a particular nucleotide is incorporated into a growing strand by DNA polymerase, the result will be a release of a hydrogen proton into solution and a subsequent change in pH level. The change in pH is detected as a voltage shift by the sensors and is recorded in real time. As the different bases (A, C, G, T) are washed sequentially through, additions are recorded, allowing the sequence from each well to be determined.
Ligation-based sequencing with 5500 Series SOLiD system exact cell chemistry
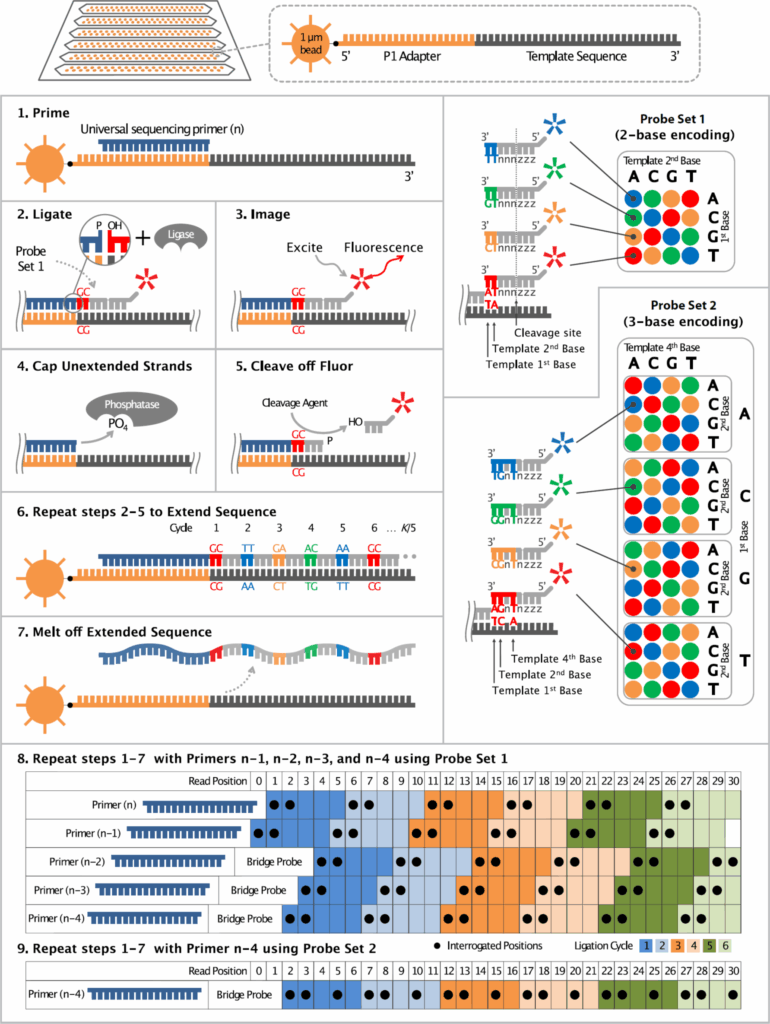
Source: Thermo Fisher Scientific.
In September 2015, Thermo launched two new NGS systems into the marketplace, Ion S5 and Ion S5 XL, which are based on the Ion Torrent technology platform. The Ion S5 and Ion S5 XL systems offer the combined ability to sequence gene panels and small genomes, exomes, and transcriptomes on a single platform. In addition, they are designed with plug-and-play, cartridge-based reagents to make setting up and operating the sequencers simple and efficient. Based on our recent discussions with potential customers at the American Society of Human Genetics Annual Meeting in Baltimore, it appears the Ion S5 and Ion S5 XL systems make targeted sequencing more accessible to academic, translational, and clinical research labs.
TABLE 10: Ion Torrent sequencers
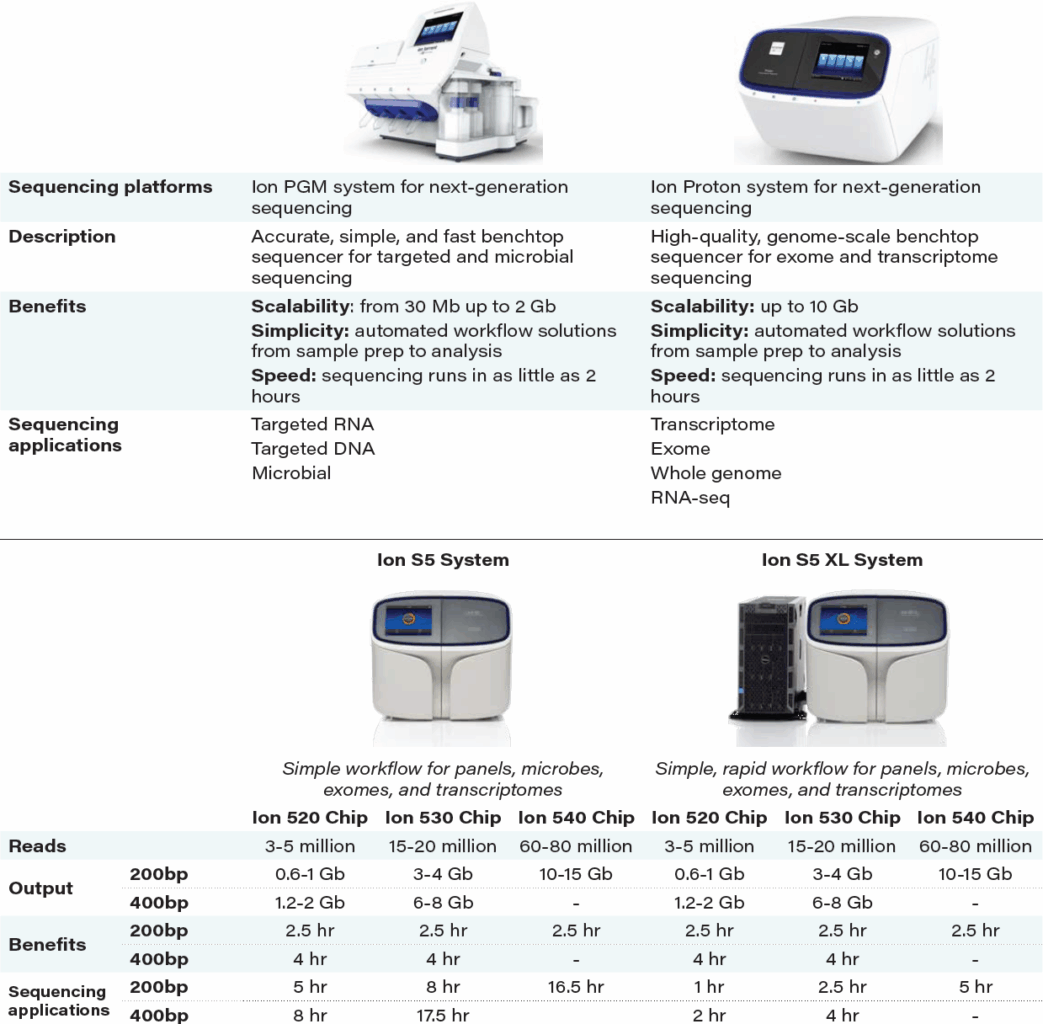
Source: Thermo Fisher Scientific.
Pacific Biosciences
Pacific Biosciences (PacBio) was founded in 2004 and went public in 2010, selling 12.5 million shares at an initial price of $16 per share. Its first commercial sequencer, the PacBio RS, was sold to a limited set of customers in 2010 and was commercially released in early 2011. A new version of the sequencer called the PacBio RS II was released in April 2013. In 2013, a partnership between Pacific Biosciences and Roche Diagnostics was announced for the development of diagnostic products for clinical use, including sequencing systems and consumables based on SMRT technology, with Roche providing $75 million in the deal.
In late 2015, the company launched a new sequencing instrument called the Sequel System with approximately sevenfold greater capacity than the PacBio RS II. PacBio’s technology uses a single-molecule, real-time sequencing (SMRT) approach. When the SMRT technology was first released, there was a great deal of concern regarding the high error rates in base calls. However, the company has since incorporated circular consensus sequencing (CCS) into its system, which has greatly reduced error rates by allowing fragments to be sequenced repeatedly to check for errors. The SMRT sequencing system involves no amplification step, setting it apart from the other major next-generation sequencing systems. The sequencing is performed on a chip containing many zero-mode waveguide (ZMW) detectors. DNA polymerases are attached to the ZMW detectors, and phospholinked dye-labeled nucleotide incorporation is imaged in real time as DNA strands are synthesized. PacBio’s RS II C2 XL currently offers both the greatest read lengths (averaging around 4,600 bases) and the highest number of reads per run (about 47,000).
SMRT sequencing with PacBio Systems

- Take advantage of the Sequel System to reduce project costs and generate 7X more reads compared with the PacBio RS II
- Achieve ~10kb average read lengths, with some reads as long as 60 kb
- Scale throughput based on project needs:
- 8-12X coverage per genome for structural variation surveys
- 25X coverage per genome for hybrid assembly
- 50X coverage per genome for PacBio-only de novo assembly
- Simultaneously capture epigenetic information
Source: Pacific Biosciences.
Oxford Nanopore
Oxford Nanopore Technologies (ONT) is a U.K.-based private company founded in 2005 as a spinout from the University of Oxford by Hagan Bayley, Gordon Sanghera, and Spike Willcocks. Since its inception, ONT has raised ~$380 million in private equity and venture funding with a goal of developing nanopore-based DNA sequencing systems for commercial use.
PromethiON

Source: Oxford Nanopore Technologies.
A nanopore is 1 nanometer in diameter and can either be simply small holes in an inorganic membrane (solid-state nanopores) or specific channels made from modified natural pore-forming proteins embedded in a membrane (biological nanopores). In the case of ONT, a protein nanopore is set in an electrically resistant polymer membrane. During sequencing, the sequencer pulls a strand of DNA through the pore and monitors the changes in the electric current to tell which of the four DNA bases is going through the opening. For example, ONT sequencing can be used to distinguish among the four standard DNA bases G, A, T, and C, and also modified bases.
Nanopore workflow
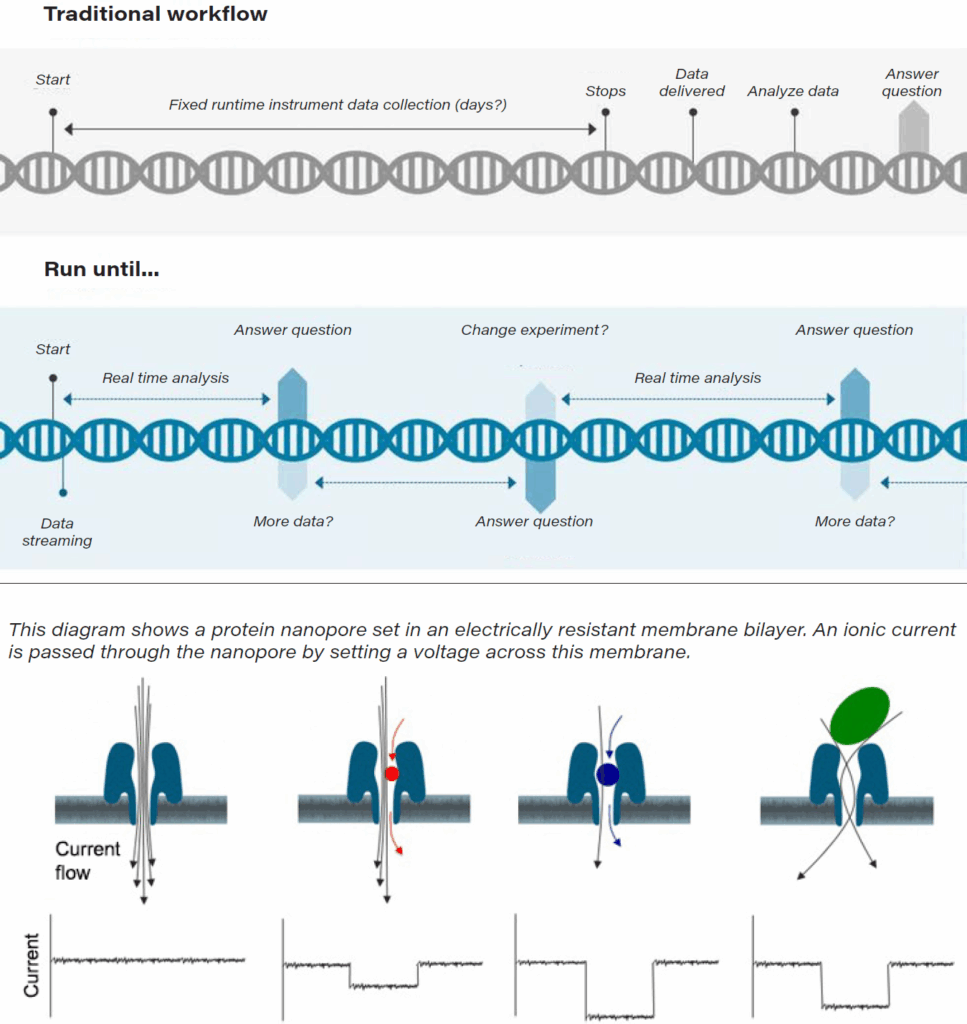
Source: Oxford Nanopore Technologies.
Nanopore
DNA can be sequenced by threading it through a microscopic pore in a membrane. Bases are identified by the way they affect ions flowing through the pore from one side of the membrane to the other.
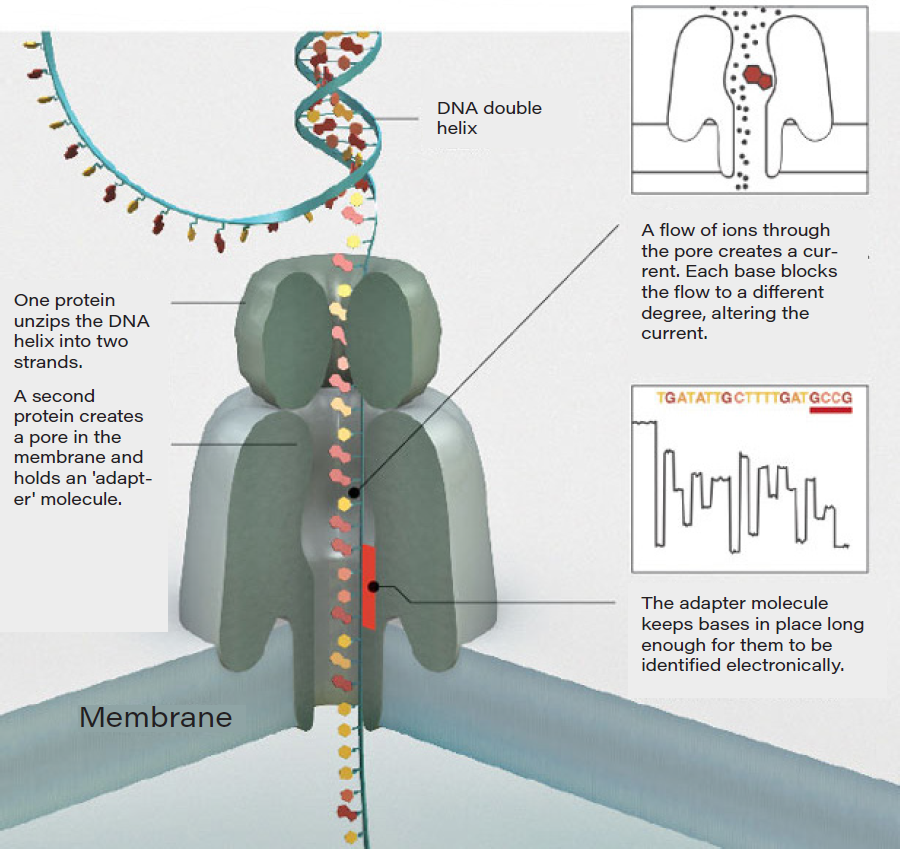
Source: Oxford Nanopore Technologies.
The advantage of this sequencing approach is that, at least in principle, any strand length of DNA can be sequenced. In addition, ONT sequencing minimizes sample preparation, eliminates the need for amplification or modification (nucleotides, polymerases, or ligases), and provides long read lengths (10,000-50,000 bases). ONT’s sequencing platform includes MinION, PromethION and GridION, all of which are adaptable for the analysis of DNA, RNA, proteins, small molecules, and other types of molecules. A key selling point of the MinION, PromethION, and GridION systems is there is no fixed run time; thus, a user can run any of the three systems for a short or long period of time as data is streamed in real time. This can enable real-time analyses so that the user can predetermine an experimental endpoint and run the system for as long as it takes to collect sufficient data. As a result, the ONT platform has a broad range of potential applications, including scientific research, healthcare, agrigenomics, and security/defense.
Although ONT sequencing has its advantages, there are significant challenges to be overcome. Among them, a high error rate of ~5-10% and the requirement of ultra-precise, high-speed DNA detection beyond the scope of existing optical and electrical technologies.
TABLE 11: Sequencing platform overview
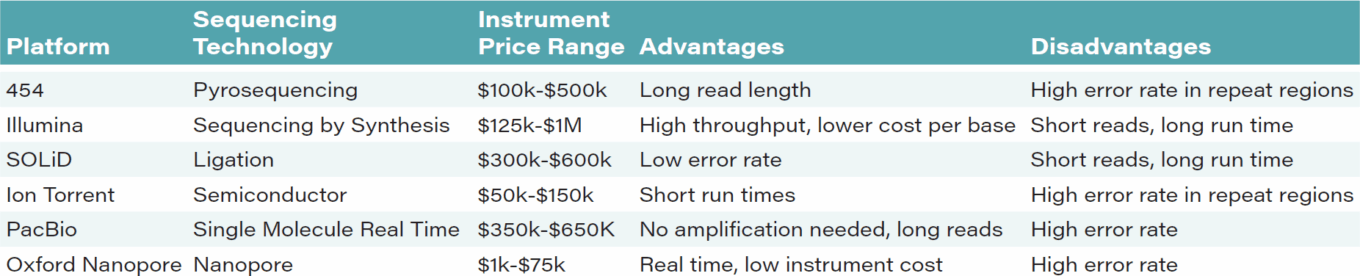
Source: First Analysis.
Key NGS focus areas
Long reads vs. short reads
Like a jigsaw puzzle with large pieces, a genome de novo sequenced with long reads is easier to assemble. Long reads enable a comprehensive view of the genome, as they can reveal multiple types of genetic variation such as structural variants. Currently, NGS technologies (with the exception of PacBio and ONT) have favored lowering per-base cost at the expense of read length. This has dramatically reduced sequencing cost but has resulted in increased challenges to alignment, or not being able to span even modest repeat regions, and genome assembly, which, in turn, can produce biases and errors when interpreting results.
Currently, short-read data (i.e., reads a few hundred bases in length) are either mapped to a reference genome or computationally assembled as a draft genome, and the structural variants are based on these mapped reads. Genome assembly via reference mapping involves aligning the reads with specific chromosomal locations on a reference genome sequence obtained from bioinformatic software and online databases. Systematic alignment of reads to incorrect positions in the genome can lead to false inferences of single-nucleotide polymorphisms and copy number variants. This is particularly problematic because single-nucleotide polymorphisms and copy number variants are viewed as the driver of mutations in many different cancers and have proven to be important diagnostic biomarkers as well as effective drug targets.
Despite the better accuracy NGS provides, it cannot make up for its inability to detect variations caused by contiguous repetitive DNA sequences. Most large genomes are filled with repetitive sequences; for example, nearly 50% of the human genome is composed of repeat regions. Repeats arise from a variety of biological mechanisms that result in extra copies of a sequence being produced and inserted into the genome. Repeats come in all shapes and sizes: They can be widely interspersed repeats, tandem repeats, or nested repeats; they may comprise just two copies or millions of copies; and they can range in size from one to two bases (mono- and dinucleotide repeats) to millions of bases. Without knowing the genomic context of these repeats or the amount of repeats, it is difficult to attach any biological significance to them; in other words, you cannot analyze what you cannot measure.
In order to bridge the gap between read lengths, Illumina acquired Moleculo in 2012 for its synthetic long-read technology and incorporated it into its TruSeq platform. The technology breaks DNA into large fragments that are sequenced on standard Illumina sequencing platforms for subsequent assembly into synthetic long reads or whole human genome phasing using proprietary informatics. According to Illumina CEO Jay Flatley, for 90% of long-read applications, synthetic long reads will suffice.
In addition to Moleculo, 10X Genomics has developed an innovative system, GemCode, providing long-range (i.e., long-read) data by upgrading existing NGS systems. The GemCode platform uses molecular barcoding to identify and tag long DNA molecules.
The GemCode System

Source: 10x Genomics.
Complementing long-range sequencing approaches are advances in long-range mapping technologies, such as BioNano Genomics’ Irys sytem, that enhance the quality of de novo genome sequencing and assemblies. Long-range mapping helps identify and label potential sequencing inconsistencies and larger forms of structural variation. The Irys system, for instance, was recently used to generate optical maps of DNA fragments ≥150 kb to assemble and resolve one of the most repetitive regions (chromosome 1q21) of the human genome.
BioNano workflow
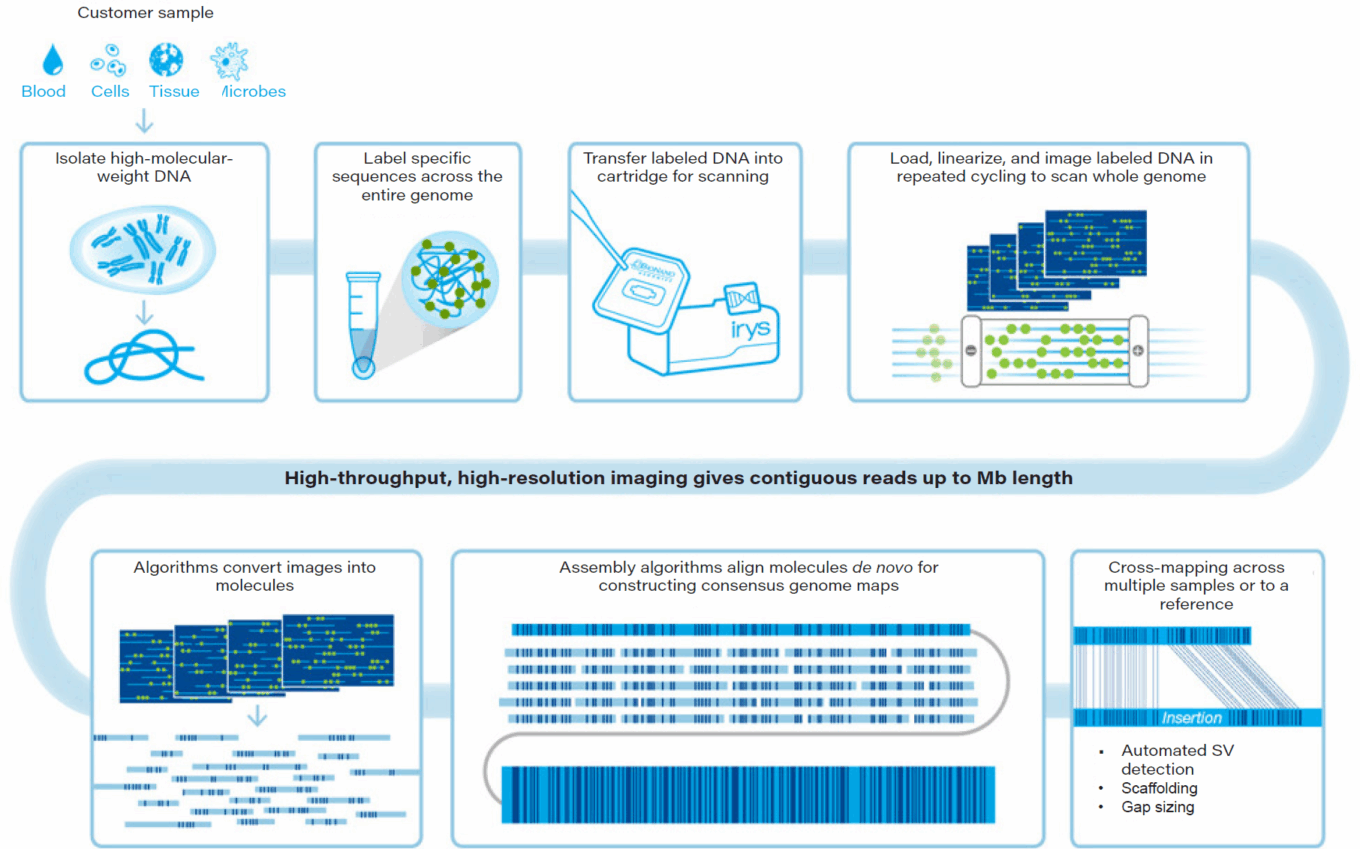
Source: BioNano Genomics.
BioNano Irys system
Source: BioNano Genomics.
Recent advances in long-read sequencing technology from PacBio, ONT, 10X Genomics, and Illumina as well as mapping technologies from BioNano have made it possible for individual laboratories to consider the possibility of generating high-quality de novo assemblies of new genomes. In our view, de novo sequencing of human genomes via long read, as opposed to alignment to a reference genome, provides a more comprehensive view and understanding of genetic variation.
Sequencing Company Profiles

10X Genomics, based in Pleasanton, Calif., is a privately held company. Through its GemCode platform, the company is focused on enhancing and upgrading the capabilities of existing short-read sequencers. It delivers additional genetic information through a combination of microfluidics, chemistry, and bioinformatics that gives researchers access to structural variants, haplotypes, and other critical genetic information.

Agilent Technologies, based in Santa Clara, Calif., is a publicly traded measurement company providing services to the bio-analytical, life sciences industries. The bio-analytical measurement business provides application-focused solutions that include instruments, software, consumables, and services that enable customers to identify, quantify, and analyze the physical and biological properties of substances and products.

BGI/Complete Genomics, based in Shenzhen, China, is a privately held genomic services company that provides comprehensive sequencing and bioinformatics services for commercial science, medical, agricultural and environmental applications. Prior to its acquisition by BGI in 2013, Complete Genomics was focused on selling whole-genome sequencing services to over 150 research customers. In 2015, BGI/Complete Genomics introduced the Revolocity system, an end-to-end genomics solution for large-scale, high-quality genomes.

BioNano Genomics, based in San Diego, Calif., is a privately held next-generation mapping (NGM) company. It provides customers with genome analysis tools that advance human, plant, and animal genomics and accelerate the development of clinical diagnostics. The company’s Irys System uses NanoChannel arrays integrated within the IrysChip to image genomes at the single-molecule level with average single-molecule lengths of about 350,000 base pairs. The long-range genomic information obtained with the Irys System helps decipher large, complex DNA repeats, which are the primary cause of inaccurate and incomplete genome assembly.

RainDance Technologies, based in Billerica, Mass., is a privately held life sciences company whose genomic tools enable research of novel noninvasive liquid biopsy applications for the early detection of cancer and other inherited and infectious diseases.
Clinical diagnostics
Molecular diagnostics is a subset of in vitro diagnostic (IVD) tests that involves the measurement of DNA, RNA, proteins, enzymes, or metabolites at the molecular level in order to detect genotypes, mutations, or biochemical changes. The core objective of molecular diagnostics is to determine whether a specific person is predisposed to have a disease, whether he/she actually has a disease, or whether a certain treatment option is likely to be effective for a specific disease.
Molecular diagnostics, though only roughly 15% of the ~$57 billion IVD industry, has emerged as one of the fastest-growing segments at ~10% annually. According to industry executives we’ve spoken with, the market is expected to continue to grow at a five-year CAGR of ~10%, driven by increasing automation, clinical adoption of NGS, expanded test panels, more genetic testing, and an expected increase in the number of labs performing molecular diagnostics tests.
TABLE 12A: 2015 molecular diagnostics market by category
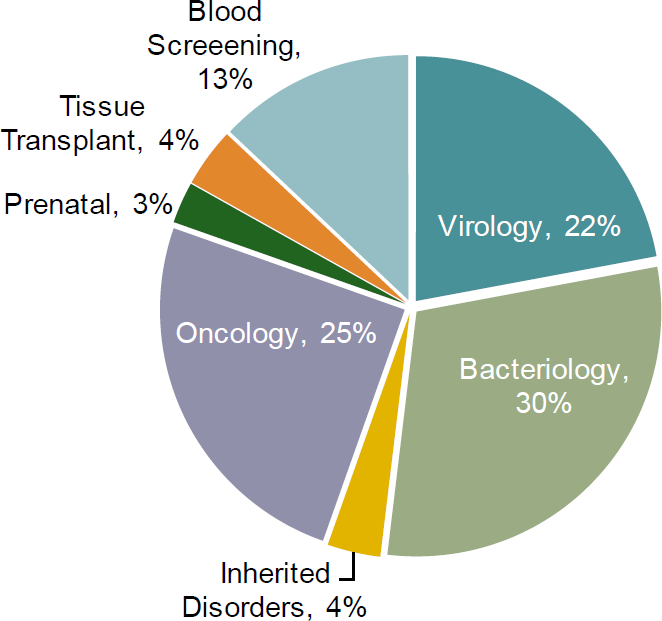
TABLE 12B: 2015 molecular diagnostics market share ~$7.7B
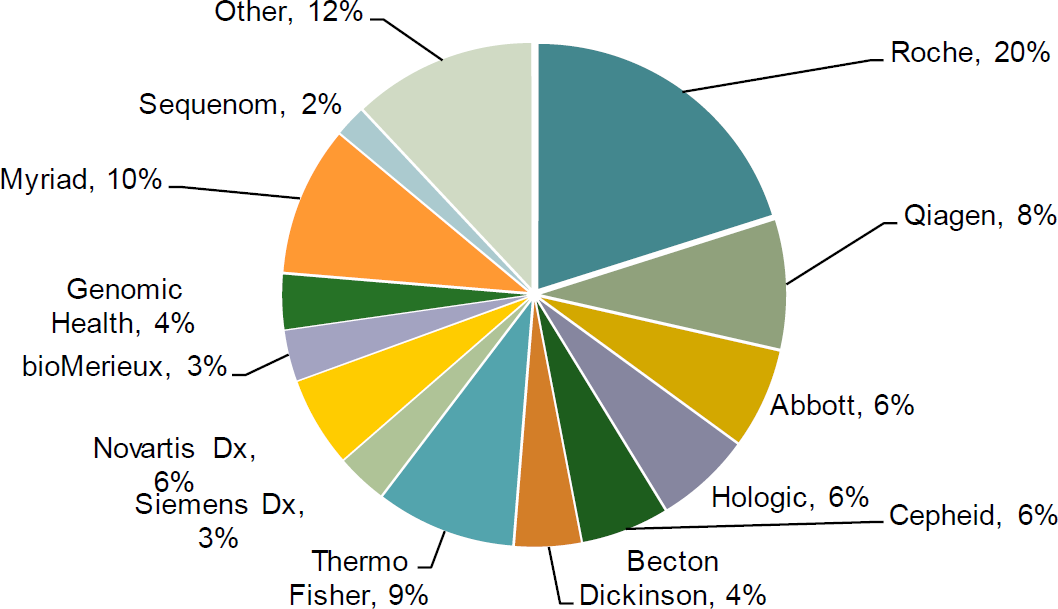
Source: First Analysis, Bloomberg, company reports.
The molecular diagnostics market consists of three broad categories of testing: clinical infectious disease, genetics, and blood screening. We estimate annual sales of test kits and reagents for molecular genetic testing are ~$2.5 billion.
- The infectious disease category includes viral (HIV, HCV, HBV, and respiratory viral testing) and bacterial (CT/NG, MRSA/SA, C. difficile, MTB, GBS, and vaginitis).
- The blood screening category includes testing of blood products such as whole blood plasma using NAT assays for HIV, HCV, HBV, WNV, HAV, and parvovirus.
- The genetics category includes tests for cancer screening, cystic fibrosis, transplantation/autoimmune (HLA), coagulation (Factor V/Factor II/MTHFR), pharmacogenetic testing (e.g., CYP450), and companion diagnostic testing (e.g., KRAS, BRAF, and EGFR).
TABLE 13: Molecular diagnostic test examples
| Test | Description | Examples |
|---|---|---|
| Newborn screening | Targeted tests for recessive genetic disorders | Cystic fibrosis, sickle-cell anemia |
| Diagnostic testing | Confirmatory test or differential diagnosis testing for a symptomatic individual | Skeletal dysplasia, craniosynostosis |
| Carrier testing | Targeted testing for asymptomatic individuals potentially carrying one or more recessive mutations | Cystic fibrosis, Tay-Sachs disease |
| Predictive testing | Tests for variants causing or associated with diseases or disorders with a hereditary component, usually with adult-onset symptoms | Cancers, cardiovascular disease, diabetes |
| Pre-symptomatic testing | Tests for variants causing or associated with diseases or disorders known to be inherited in the family, often with adult-onset symptoms | Huntington’s disease, Alzheimer’s disease |
| Pharmacogenetics | Targeted tests for variants associated with pharmaceutical dosage choice or adverse reactions | DNA test for warfarin |
Source: First Analysis, Bloomberg, company reports.
Illustrative growth scenarios for molecular diagnostic and genetic testing spending, 2010-2021
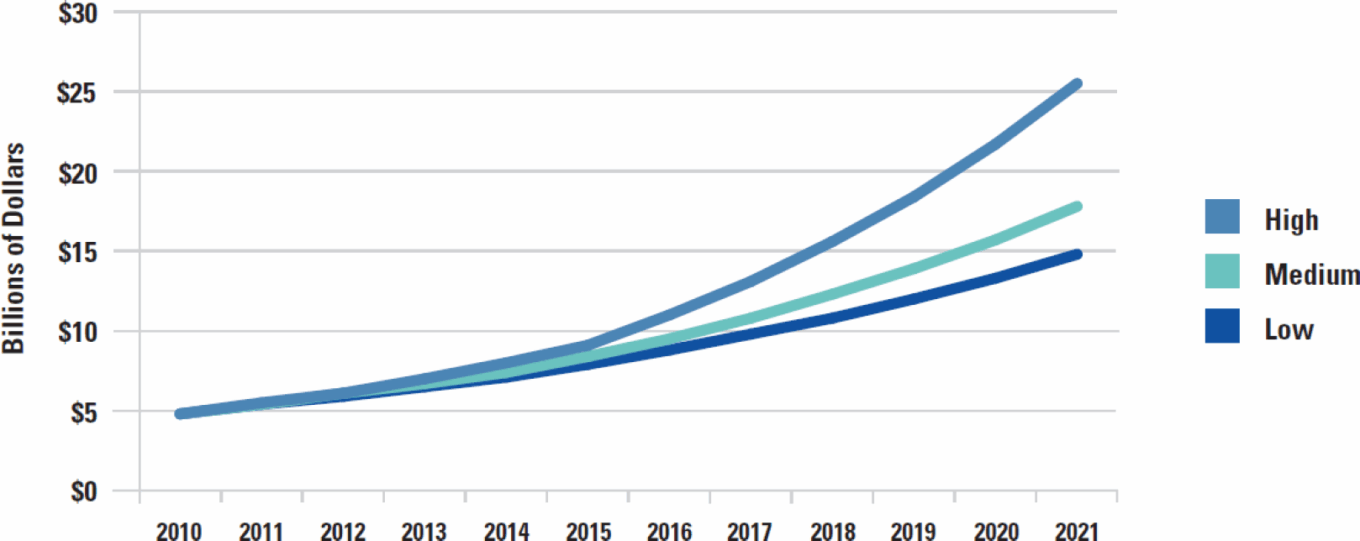
Source: United Health.
NGS in molecular diagnostics
NGS tests are an important and growing part of the molecular diagnostics market. Unlike other laboratory tests that typically detect a single or a defined number of substances to diagnose a limited set of conditions, a single NGS test can identify thousands—even millions—of genetic variants, and the results of that test could be used to diagnose or predict an individual’s risk of developing many different conditions or diseases. If it were possible to accurately and cost-effectively sequence the whole genome, it would not be necessary to know what variant one wishes to identify prior to running and successfully interpreting an NGS test—a concept that is very different from how traditional IVDs are used.
What is an ‘NGS’ test?
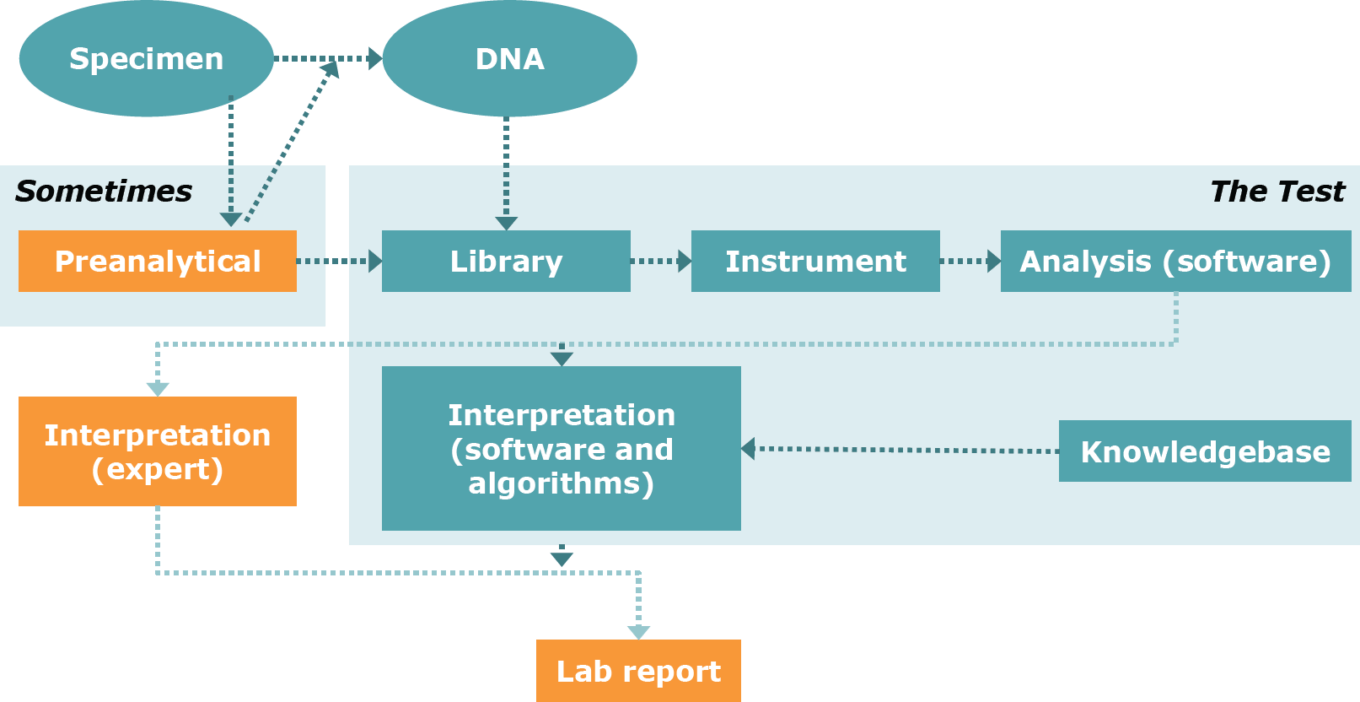
Source: FDA.
As promising as the NGS technology is, many issues remain to be resolved before widespread clinical adoption. These include issues around bioinformatics and results interpretation, cost, reimbursement, informed consent, assay validation, reference materials, and quality control.
TABLE 14: Approved personalized medicine oncology drugs as of 20141
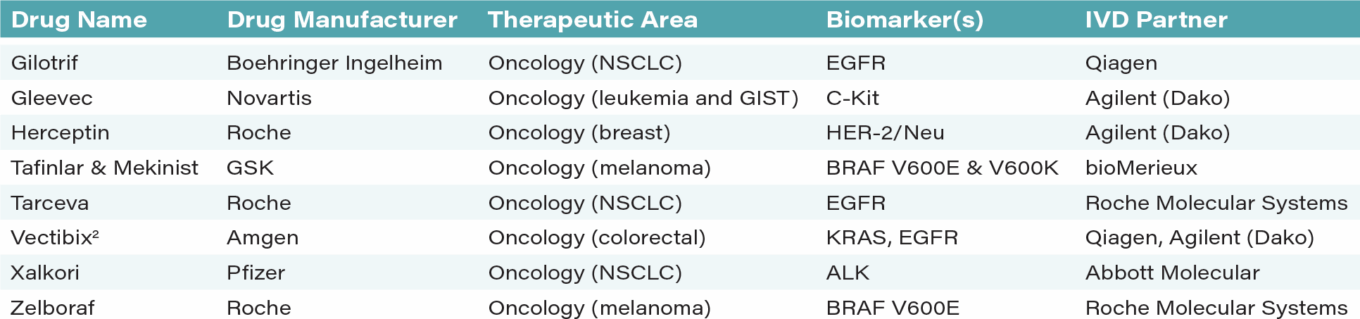
Source: L.E.K. Consulting, First Analysis.
Notes: (1) Based on List of Cleared or Approved Companion Diagnostic Devices by FDA in December 2014. (2) Similar tests are approved for Erbitux.
Companion diagnostics
According to the FDA guidance document issued in August 2014, a companion diagnostic is defined as an IVD device that provides information essential for the safe and effective use of a corresponding therapeutic product. Furthermore, the FDA states companion diagnostics “identify patients who are most likely to benefit from a particular therapeutic product” or are “likely to be at increased risk for serious adverse reactions as a result of treatment with a particular therapeutic product.” The labeling instructions of the therapeutic product would stipulate the use of the IVD companion diagnostic device. Despite much ongoing research within chronic diseases such as autoimmune disorders and neurological and cardiovascular diseases, companion diagnostics is still mainly related to oncology. Typically, most pharmaceutical companies lack internal diagnostic capabilities for companion diagnostics testing, so they partner with an IVD manufacturer for companion diagnostics development and regulatory submission.
TABLE 15A: Worldwide companion-diagnostics-informed drug revenue 2013*
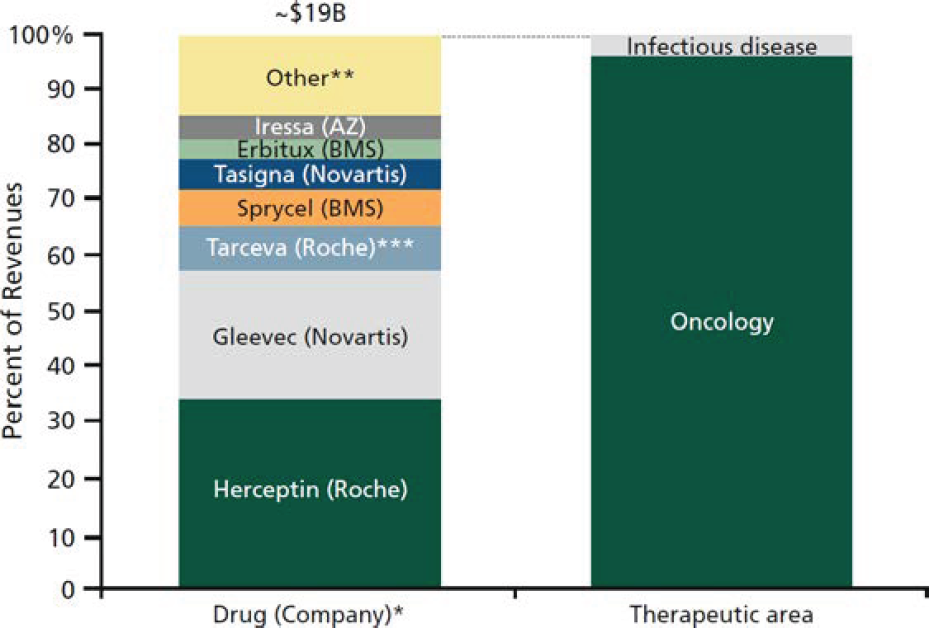
TABLE 15B: Late-stage clinical trial pipeline by trial type and therapeutic area

Source: L.E.K. Consulting.
Notes: *2013 revenues are actual or analyst estimates; products include those with labels that require/recommend companion diagnostics tests for candidacy. **“Other” includes Mekinist, Bosulif, Tafinlar, Vectibix, Selzentry, Kadcyla, Xalkori, Tykerb/Tyverb, Perjeta, Zelboraf, and Victrelis. ***Includes all Tarceva revenues.
Pharmacogenetics
Another emerging area within the field of molecular diagnostics is pharmacogenetics testing. This refers to testing a patient for a gene mutation that may affect his or her ability to metabolize or respond to a drug, which may result in toxicity or lack of efficacy. When sufficient clinical information is available, pharmacogenetics may also aid in dosage selection of the therapeutic. Although the primary focus of pharmacogenetics testing has been on improving drug selection and dosing in patient populations or individuals, a secondary potential benefit of testing may be the improvement of medication adherence. Medication adherence is a well-documented problem in the United States, with annual costs tied to nonadherence of ~$300 billion.
TABLE 16: Therapeutic impact of pharmacogenetics
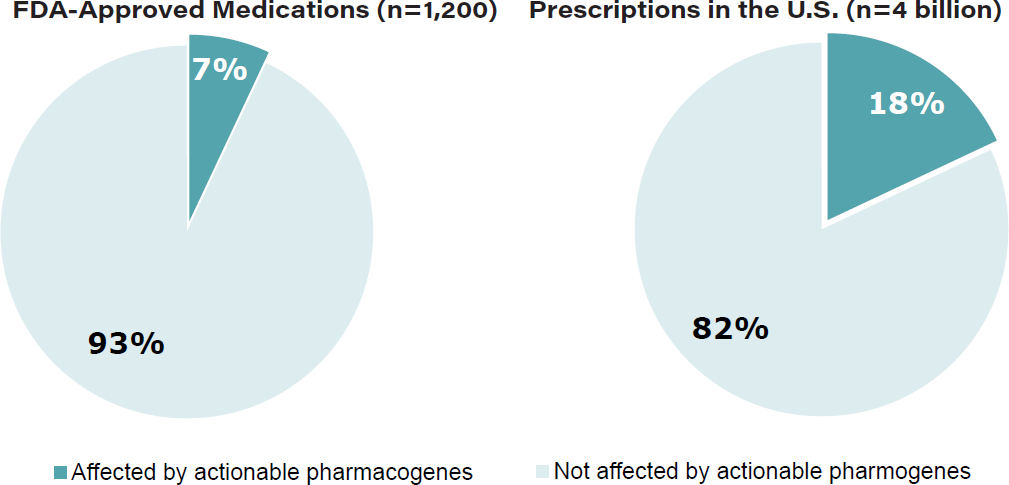
Source: FDA, company reports.
TABLE 17: Pharmacogenetics examples
| Genetic variation | Medications |
|---|---|
| TPMT | Mercaptopurine, thioguanine, azathioprine |
| CYP2D6 | Codeine, tramadol, tricyclic antidepressants |
| CYP2C19 | Tricycle antidepressants, clopidogrel, voriconazole |
| VK0RC1 | Warfarin |
| CYP2C9 | Warfarin, phenytoin |
| HLA-B | Allopurinol, carbamazepine, abacavir, phenytoin |
| CFTR | Ivacaftor |
| DPYD | Fluorouracil, capecitabine, tegafur |
| G6PD | Rasburicase |
| UGT1A1 | Irinotecan, atazanavir |
| SLC01B1 | Simvastatin |
| IFNL3 (IL28B) | Interferon |
| CYP3A5 | Tacrolimus |
Source: FDA, company reports.
Although the potential benefit of pharmacogenetics testing to improve adherence has been exhibited in chronic diseases such as diabetes, we envision pharmacogenetics as a future intervention tool, either alone or in combination with other modalities.
Barriers to pharmacogenetics adoption
The barriers to adoption of pharmacogenetics among clinicians include the following:
- The limited knowledge of pharmacogenetics among physicians makes the results of the tests difficult to translate into clinical decisions.
- As an emerging field of scientific discovery impacting the practice of medicine, widespread adoption of pharmacogenetics has been slow, pending further evidence supporting its efficacy.
- The complexity of results creates a negative predisposition among clinicians to the usefulness of pharmacogenetics.
- Adoption of testing has been slow due to reimbursement inconsistencies among payers.
- Physicians are unsure which patients to test.
- Cost.
Liquid biopsy
The concept of liquid biopsy typically revolves around the idea that DNA from cancer cells is present within the bloodstream, not just in the tissue of origin, and can be detected from a blood test. The noninvasive test involves capturing and analyzing circulating tumor cells (CTC) in the blood stream as well as cell-free DNA (cfDNA) shed from tumors. The CTC or cfDNA, once isolated, can then be analyzed to better understand the molecular profile of the tumor, allowing the physician to choose the best course of treatment. In theory, a liquid biopsy could be done at any time to determine if patients are responding to treatments or if there has been disease recurrence. Although liquid biopsy is not quite ready for prime time today and not likely to be for at least three to five years, we estimate the potential market opportunity to be ~$15 billion with numerous companies working on technologies and solutions.
Laboratory-developed tests
Traditionally, most genetic tests have not been subject to premarket review by the FDA. This is because in the past, genetic tests were developed by laboratories primarily for their in-house use—referred to as laboratory-developed tests—to diagnose mostly rare diseases and were highly dependent on expert interpretation. However, more recently, laboratory-developed tests have been developed to assess relatively common diseases and conditions, thus affecting more people, and direct-to-consumer (DTC) genetic testing has become more available over the Internet.
Laboratory-developed tests are defined as diagnostic tests developed and used within a single lab. Laboratory-developed tests also go by the nickname “home brews” due to the fact that many large molecular laboratories have developed the skill sets to make their own laboratory-developed tests. The laboratories that perform these tests are subject to the Clinical Laboratory Improvement Amendment (CLIA) rules, administered and implemented by the Centers for Medicare and Medicaid Services (CMS).
Laboratory-developed tests
Diagnostic tests are developed either by manufacturers for distribution to laboratories or by laboratories themselves for use in their facilities. The tests developed by labs are referred to as laboratory-developed tests (LDTs).
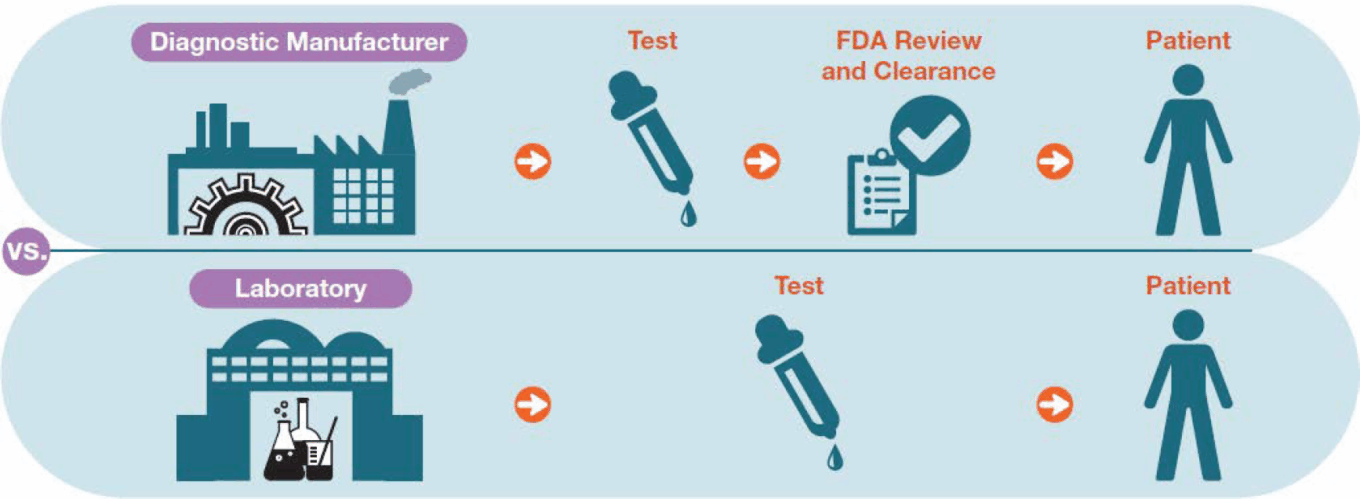
Source: AdvaMedDx..
Clinical labs can obtain CLIA certification directly from CMS, typically through state agencies that survey labs for compliance with CLIA requirements. In addition, certification can occur if a lab is accredited by one of the independent accreditation organizations approved by CMS. These include the College of American Pathologists (CAP) and the Commission on Office Laboratory Accreditation (COLA), among others. Before approving an independent accreditation organization, CMS must determine the organization’s standards are equal to or more stringent than those set forth in the CLIA regulations, though the standards may differ from CLIA by including additional requirements. Certification can be issued if a lab is accredited by one of the independent accreditation organizations approved by CMS.
The National Institutes of Health’s Genetic Testing Registry (GTR) currently contains more than 32,000 tests from nearly 450 labs. The recent explosion of NGS-based test offerings has led the FDA to publish guidance documents on the regulation of these products. Traditionally, diagnostic tests have fallen into two main categories, in vitro diagnostics and laboratory-developed tests. The former are products containing all the reagents and materials needed to run the test and are regulated by the FDA as medical devices. Today, most NGS-based clinical tests are classified as laboratory-developed tests, with two exceptions: In 2013, the FDA approved Illumina’s cystic fibrosis carrier screening assay, an assay that detects 139 variants in the cystic fibrosis transmembrane conductance regulator (CFTR) gene, and Illumina’s assay for CF diagnosis by sequencing all the medically relevant regions of the CFTR gene.
Although the FDA has long regulated in vitro diagnostic products as medical devices and has taken the position it has the authority to regulate laboratory-developed tests, the agency historically has exercised “enforcement discretion” and has not actively regulated laboratory-developed tests. In November 2015, the FDA announced it would move forward with its 2014 draft guidance and finalize a plan to regulate laboratory-developed tests as IVDs sometime in 2016 under the federal Food, Drug, and Cosmetic Act.
The FDA’s 2014 draft guidance states the agency will use a risk-based approach to regulating laboratory-developed tests. The FDA will rely upon the existing medical device classification system to evaluate the risk of a category of laboratory-developed tests. Medical devices are classified as Class I, II, or III based on the controls necessary to provide a reasonable assurance of the safety and effectiveness of the device, and factors relevant to this determination include the device’s intended use, technological characteristics, and the risk to patients should the device fail.
Class I devices, which are subject only to general controls, typically represent the lowest-risk category of devices, while Class III devices, which are subject to general controls and premarket approval, often represent the highest-risk devices. In determining the risk an LDT poses to the patient and/or the user, the FDA will consider a series of factors, including whether the device is intended for use in high-risk diseases/conditions or patient populations, whether the device is used for screening or diagnosis, the nature of the clinical decision that will be made based on the test result, whether a physician/pathologist would have other information about the patient to assist in making a clinical decision (in addition to the LDT result), alternative diagnostic and treatment options available to the patient, the potential consequences/impact of erroneous results, and the number and type of adverse events associated with the test, among others.
To provide additional clarity, the FDA intends to formalize what it considers generally to be Class I, II, or III, and the agency has proposed to phase in the review process over 10 years, with high-risk laboratory-developed tests being reviewed first, followed by moderate-risk ones.
TABLE 18: LDT oversight framework summary: requirements FDA intends to enforce
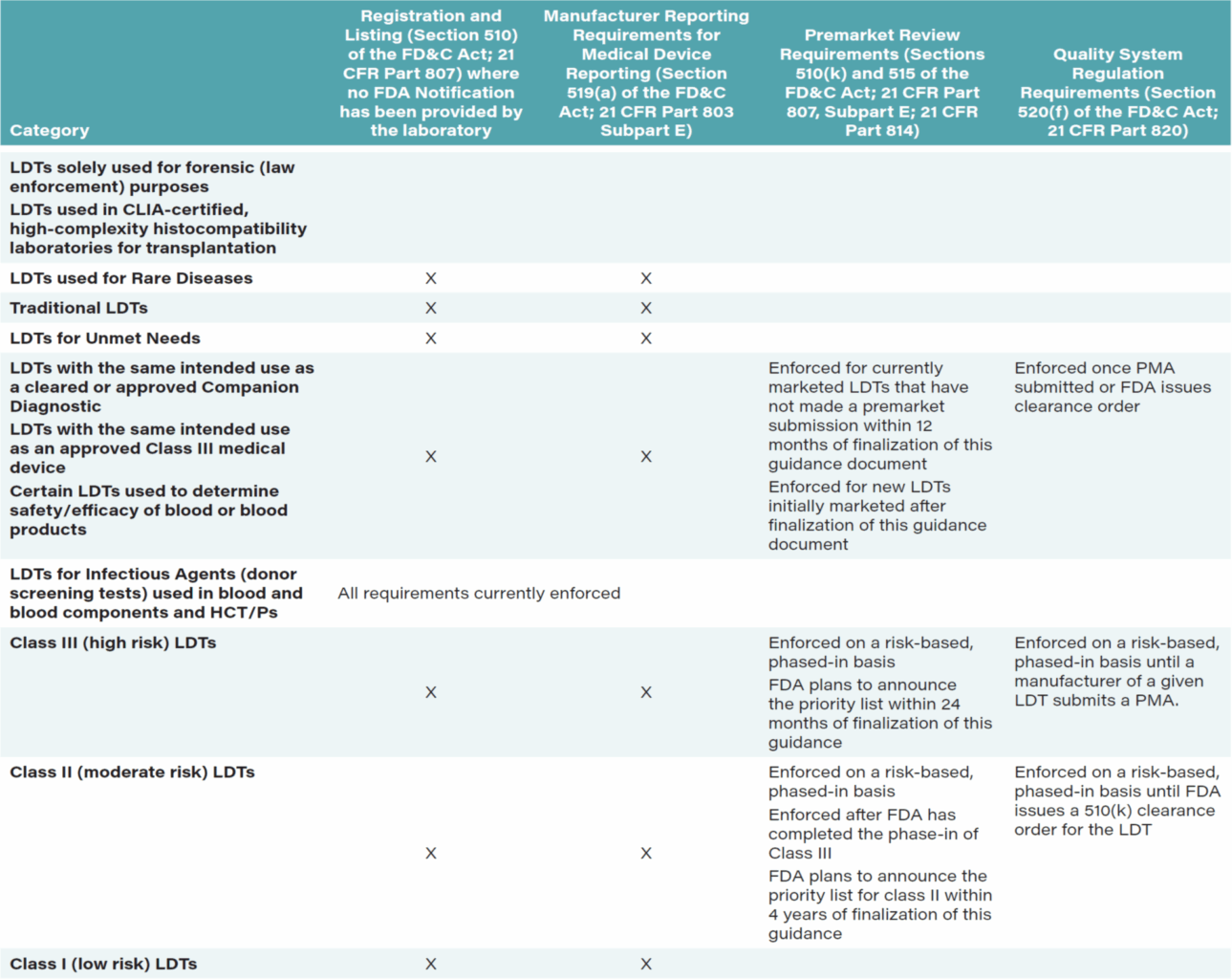
Source: FDA.
Some thoughts on data interpretation, bioinformatics
Currently, the ability to collect massive amounts of genetic data outpaces the ability to understand and act on it, and while the overall cost of sequencing is falling at a dramatic rate, the costs and issues associated with interpreting, storing, and managing the data are steadily increasing. In addition, the current lack of industry standards for both accuracy and quality has only exacerbated the questions surrounding clinical utility and validity.
In our view, the data must be cost-effectively managed so that the clinically relevant and validated information can be extracted, analyzed, conveyed, and stored in an understandable way for clinicians. In addition, automation traceability and privacy workflow solutions are needed for successful integration with electronic medical records (EMRs).
Reimbursement dynamics
Clinical laboratories face significant coding and reimbursement challenges in 2016, especially for NGS. As testing technologies advance, clinical applications rapidly evolve, and utilization increases, a patchwork billing and payment framework is being forced to evolve and change.
Prior to 2013 and an inflection point in the perceived utility and adoption of genetic testing, coding for clinical lab tests that involved DNA or RNA measurement was typically done via “stacked codes,” or generic process codes describing specific steps (see examples in Table 20), and payment was determined by summing the payment values of the steps coded. In 2013, the proliferation of molecular diagnostics and increased confidence in its utility led CMS to establish an index of specific codes. The switch led to lower reimbursement rates and a substantial amount of confusion, which in turn led to an almost complete lack of federal reimbursement in the first quarter of 2013.
TABLE 19: Evaluating the validity of genetic tests
| Term | Definition |
|---|---|
| Analytical sensitivity | The proportion of assays with the genotype that have a positive test result. |
| Analytical specificity | The proportion of assays without the genotype that have a negative test result. |
| Clinical sensitivity | Refers to the proportion of people with a disease who have a positive test result. |
| Clinical specificity | Refers to the proportion of people without a disease who have a negative test result. |
| Positive predictive value | Refers to the likelihood that a patient has the disease given the test result is positive. |
| Negative predictive value | Refers to the likelihood that a patient does not have the disease given the test result is negative. |
| Clinical utility | Refers to the value of the test for determining treatment, patient management, and family planning. |
| Personal utility | Refers to the value of the test for personal and family choices. |
Source: First Analysis.
Now, CMS is undertaking the same process in regard to NGS test coding and reimbursement. In September 2015, CMS announced it would implement the Protecting Access to Medicare Act of 2014 (PAMA). PAMA mandates a completely new market-based Medicare payment schedule for clinical laboratory tests. Under PAMA, clinical laboratories and physician offices are required to report the volume and fee amounts received from private insurers if they have more than $50,000 in Medicare revenue from laboratory services and they receive more than 50 percent of their Medicare revenue from laboratory and physician services. Laboratories would collect private payer data from July 1, 2015, through Dec. 31, 2015, and report it to CMS by March 31, 2016. CMS will post the new Medicare Clinical Laboratory Fee Schedule (MCLFS) rates (based on weighted median private payer rates) in November 2016, which will be effective on Jan. 1, 2017.
TABLE 20: Examples of genetic testing billing codes used before and after 2013 (Medicare Released AMA Molecular Test Codes)
| Before 2013 | From 2013 forward |
|---|---|
| “Stack codes” for DNA/RNA processes | Gene-specific codes (“Tier 1”) |
| 83898 DNA Amplification Step 83904 DNA Sequencing Step 83912 Sequence Report and Interpretation | 81200 ASPA (aspartoacylase, Canavan disease) gene analysis 81220 CFTR (cystic fibrosis gene), common variants 81310 NPM1 (nucleoplasmin, acute myeloid leukemia), exon 12 variants |
| ~ 500 genes classified in “Tier 2, Levels 1-9” | |
| 81400 Level 1 Molecular Procedure, e.g., single germline variant, single nucleotide polymorphism 82404 Level 4 Molecular Procedure, e.g., analysis of single gene exon or >10 amplicons Each “level” may list up to dozens of genes to be represented on insurance claims by a single level code, e.g., 82404 |
Source: Precision Medicine Coalition, First Analysis.
PAMA provides laboratories with some protection from short-term payment reductions by limiting any annual Medicare reduction due to market-based pricing to 10% in 2017- 2019 and to 15% in 2020-2022. As of now, after 2022 there are no further reduction limits.
TABLE 21: Examples of NGS test costs to be considered in new CMS fee schedule
| Genomic sequencing procedure | Average cost | CPT code(s) |
|---|---|---|
| <50 Genes Tumor | $691.07 | 81445, 81455 |
| Targeted Genetics Panel | $1,450.35 | 81430, 81470 |
| Whole Exome | $2,404.74 | 81415, 81416, 81417 |
Source: First Analysis, Association of Molecular Pathology.
PAMA addresses the implementation of changes to the MCLFS for Advanced Diagnostic Laboratory Tests (ADLTs), and the definition of an ADLT is written to be broadly encompassing. An ADLT is defined as a clinical laboratory test that 1) is covered under Medicare Part B that is marketed and performed by a single laboratory and not sold for use by another laboratory that is either cleared or approved by the FDA or 2) meets the following criteria:
- The test is a molecular pathology analysis of multiple biomarkers of DNA or RNA.
- When combined with an empirically derived algorithm, the test yields a result that predicts the probability a specific individual patient will develop a certain condition(s) or respond to a particular therapy.
- The test provides new clinical diagnostic information that cannot be obtained from any other test or combination of tests.
- The test may include other assays.
New ADLTs for which payment has not been made under the MCLFS prior to Jan. 1, 2017, and that meet the criteria for being considered new advanced tests will be paid at actual list charge for the first three quarters the ADLT is on the market. Once the nine-month period is over, payment for new advanced tests would be based on the weighted median private payer rate reported by the single laboratory that performs the new ADLT.
In addition to PAMA, we see the transition to ICD-10 affecting test reimbursement in the short term. The replacement of ICD-9 diagnosis codes by ICD-10, which became effective on Oct. 1, 2015, has led to much confusion among payers and clinical labs. Coding errors can have negative impacts on a clinical lab, and the failure to update billing systems to reflect test coding changes can result in rejection of claims, resubmissions, increased billing costs, and delays in payment. Failure to provide valid and appropriate diagnosis codes in billing can have the same impact.
Currently, the billing code used to document the clinical appropriateness of a laboratory test will need to be taken from the new 69,000 seven-digit ICD-10 diagnosis codes. The past 14,000 five-digit ICD-9 diagnosis codes are now invalid for billing purposes, and use of these codes when coverage requires diagnostic justification will result in claim rejections.
Data interpretation/bioinformatics and molecular diagnostic testing company profiles

Ambry Genetics, based in Aliso Viejo, Calif., is a privately held genetic services company specializing in clinical diagnostics and has a comprehensive testing menu. The company’s lab in Orange County, Calif., is CAP-accredited and CLIA-certified. The company prides itself on its reputation for sharing data while safeguarding patient privacy, unparalleled service, and responsibly applying new technologies to the clinical molecular diagnostics market.

Assurex Health, based in Mason, Ohio, is a privately held personalized medicine company that specializes in pharmacogenomics. Assurex is focused on helping healthcare providers get the genetic information they need to determine the genetically appropriate medication(s) for patients suffering from neuropsychiatric and other medical conditions. Assurex’s proprietary technology is based on pharmacogenomics—the study of the genetic factors that influence an individual’s response to drug treatments—as well as evidence-based medicine and clinical pharmacology. The company’s flagship offering, GeneSight, is a genetic test developed in the company’s own clinical laboratory and is based on patented technology licensed from Mayo Clinic and Cincinnati Children’s Hospital Medical Center.

bioMérieux, based in Marcy-l’Étoile, France, is a publicly traded company that provides diagnostic solutions (reagents, instruments, software) that determine the source of disease and contamination to improve patient health and ensure consumer safety. Its products are used for diagnosing infectious diseases and providing high medical value results for cancer screening and monitoring and cardiovascular emergencies. They are also used for detecting microorganisms in agriculture, pharmaceutical, and cosmetic products.

Blueprint Genetics, based in Helsinki, Finland and San Francisco, Calif., is a privately held genetics company led by a team of cardiologists, geneticists, bioinformaticians, DNA biologists, and business developers that provides comprehensive and high-quality genetic diagnostics with next-generation sequencing. Blueprint uses a unique targeted sequencing technology called OS-Seq, which was originally developed at Stanford University. The company prides itself on providing superior genetic diagnostics and high-quality clinical interpretation as well as fast lead times and cost efficiency.

Cepheid, based in Sunnyvale, Calif., is a publicly traded molecular diagnostics company that develops, manufactures, and markets fully integrated systems for testing in the clinical market and applications in the nonclinical market. Its systems enable rapid, sophisticated molecular testing for organisms and genetic-based diseases by automating otherwise complex manual laboratory procedures. The company is committed to improving healthcare with its GeneXpert system and its expanding portfolio of Xpert tests, which span healthcare-associated infections and other infectious diseases, women’s health, and oncology.

Coriell Life Sciences (CLS), based in Camden, N.J., has developed a genomic data ecosystem that offers storage of genomic data, expert interpretation, and an interchange framework that delivers clinically relevant genomic interpretation at the point of care. Anchored by an ongoing partnership with the Coriell Institute for Medical Research and IBM, CLS was formed to scale up the robust framework and process architecture developed for the Coriell Personalized Medicine Collaborative. The company’s core products – GeneVault, GeneExchange, and GeneDose – integrate genomic medicine into clinical care while managing the breadth of information that is obtained through genomic sequencing.

Courtagen, based in Woburn, Mass., is a privately held life sciences and molecular information company that converts genomic data into actionable clinical information for the diagnosis of critical pediatric neurological and metabolic disorders. Specifically, Courtagen focuses on mitochondrial disorders, epilepsy, and intellectual disability, including autism spectrum disorders. Courtagen’s state-of-the-art next-generation sequencing clinical laboratory integrates genotype, phenotype, and disease mechanism data using cloud-based computing and custom analytical methods to provide comprehensive results for clinicians, patients, and their families to better understand and treat underlying diseases. First Analysis Corp. is a venture capital investor in Courtagen.

Foundation Medicine, based in Cambridge, Mass., is a publicly traded molecular information company (Roche 56% ownership stake) that develops, manufactures, and sells genomic analysis diagnostics for solid and circulating cancers. Its tests are based on next-generation sequencing technology. The company’s flagship offerings, FoundationOne for solid tumors and FoundationOne Heme for hematologic malignancies and sarcomas, provide a comprehensive genomic profile to identify the molecular alterations in a patient’s cancer and match them with relevant targeted therapies and clinical trials. Foundation Medicine’s molecular information platform aims to improve day-to-day care for patients by serving the needs of clinicians, academic researchers, and drug developers to help advance the science of molecular medicine in cancer.

GenomOncology, based in Cleveland, Ohio, is a genomics technology and services company. It focuses on enabling precision medicine by translating next-generation sequencing data into actionable information for both clinicians and researchers. Its GO Precision Medicine Portfolio is a suite of products and services trusted and used by molecular pathologists and medical geneticists for test validation and production, clinical decision support, and analytics for both monitoring and discovery. In addition, GenomOncology’s technology allows multiple modes of genetic analysis to fuel integrated clinical decision support and research built on accumulating experience.

Good Start Genetics, based in Cambridge, Mass., is a privately held molecular genetics information company focused in the area of reproductive medicine. Its suite of reproductive genetics products provides clinicians and patients with insightful and actionable information in order to promote successful pregnancies and healthy families. Its flagship genetic carrier screening service, GeneVu, is a comprehensive menu of highly accurate tests for known and novel mutations that cause inherited genetic disorders, and its advanced preimplantation genetic screening test, EmbryVu, is helping a wider range of couples find their paths to pregnancy at significantly lower costs. The company complements these tests and its proprietary next-generation DNA sequencing capabilities with customer care and genetic counseling to better help expecting families.

Invitae, based in San Francisco, Calif., is a publicly traded genetic information company that provides genetic diagnostics for various hereditary disorders. The company currently provides a single diagnostic service comprising hundreds of genes for a variety of genetic disorders associated with oncology, cardiology, neurology, pediatrics, and other rare disease areas. Its Family History Tool enables users to digitally build, modify, share, and save patient pedigrees as well as assess their risks and decide on the appropriate genetic test. Clinvitae is a database of clinically observed genetic variants aggregated from public sources. The company was incorporated in 2010 as Locus Development Inc. and changed its name to Invitae Corp. in 2012.

Metabolon, based in Durham, N.C., is a privately held company focused on metabolomics, the systematic study of the unique chemical fingerprints in a biological cell, tissue, organ, or organism, which are the end products of cellular processes. The company’s proprietary platforms and informatics are delivering biomarker discoveries, innovative diagnostic tests, advances in precision medicine, and meaningful partnerships in genomics-based health initiatives. Metabolon’s expertise is also accelerating research and product development across the pharmaceutical, biotechnology, consumer products, agriculture, and nutrition industries as well as academic and government organizations.

Millennium Health, based in San Diego, Calif., is a privately held healthcare solutions company aiming to deliver accurate, timely, clinically actionable information for treatment decisions. In order for patients to have the safest and most-effective treatments, the company provides clinicians and payers with personalized medical intelligence through a comprehensive and robust suite of services, including RxAnte’s population drug therapy management platforms, Millennium Pharmacogenetic Testing (PGT), and Millennium Urine Drug Testing (UDT), which can be used to better tailor patient care.

NantHealth, based in Culver City, Calif., is a subsidiary of NantWorks, a company founded and led by Dr. Patrick Soon-Shiong. NantHealth is a cloud-based healthcare IT company converging biomolecular medicine and bioinformatics with technology services through a single integrated clinical platform, providing actionable health information at the point of care, in the time of need, anywhere, anytime. By converging molecular science, near-real-time patient signal monitoring, computer science, and big data technology, the NantHealth Operating System (NantOS) platform empowers providers, patients, and payers to coordinate best care, monitor outcomes, and control cost in real time. NantHealth’s solutions enable real-time data capture from multiple disparate data sources, allowing for coordinated medical care at a lower cost.

Natera, based in San Carlos, Calif., is a publicly traded genetic testing company that operates a CLIA-certified laboratory. It specializes in analyzing microscopic quantities of DNA for reproductive health indications to provide preconception and prenatal genetic testing services primarily to OB/GYN physicians and in vitro fertilization centers. In early 2013, the company launched Panorama, a noninvasive prenatal test for pregnant women that screens for the most common chromosomal anomalies in a fetus as early as nine weeks of gestation. Other services include tests for preimplantation genetic diagnosis and miscarriage testing to determine the cause of a pregnancy loss. A noninvasive paternity test based on Natera’s technology was brought to market in August 2011 through a partnership with DNA Diagnostics Center (DDC), which holds a license to the technology in the United States.

NeoGenomics, based in Fort Myers, Fla., is a publicly traded cancer genetics testing company that operates a network of CLIA-certified clinical laboratories. The company’s testing services include cytogenetics, fluorescence in-situ hybridization (FISH), flow cytometry, immunohistochemistry, anatomic pathology, and molecular genetic testing. NeoGenomics services the needs of pathologists, oncologists, other clinicians, and hospitals throughout the United States and has laboratories in Nashville, Tenn; Irvine, Fresno, and West Sacramento, Calif.; and Tampa and Fort Myers, Fla.

OPKO Health, based in Miami, Fla., is a publicly traded diversified healthcare company. Its diagnostic testing business includes Bio-Reference Laboratories, the nation’s third-largest clinical laboratory with core genetic testing and a 420-person salesforce. New test offerings include the 4Kscore prostate cancer test and the Claros1 in-office immunoassay platform. Its pharmaceutical and biologics business features Rayaldee for secondary hyperparathyroidism in stage 3-4 chronic kidney disease patients with vitamin D deficiency; Varubi for chemotherapy-induced nausea and vomiting; hGH-CTP, a once-weekly human growth hormone injection (in collaboration with Pfizer); and Factor VIIa for treating hemophilia.

Qiagen, based in Venlo, Netherlands and Hilden, Germany, is a publicly traded provider of molecular sample and assay technologies for molecular diagnostics, applied testing, and academic and pharmaceutical research. Sample technologies are used to collect samples of tissue, fluids, etc., and stabilize, extract, and purify various molecules of interest such as DNA, RNA, or proteins from other cellular components. Assay technologies are subsequently used to amplify and enrich this small amount of isolated material to make it visible, readable, and ready for interpretation.

Roche Diagnostics, based in Basel, Switzerland, is a diagnostic division of publicly traded Hoffmann-La Roche, which manufactures equipment and reagents for research and medical diagnostic applications. Roche is the global leader in in vitro diagnostics and tissue-based cancer diagnostics. Roche’s personalized healthcare strategy is focused on providing medicines and diagnostics that enable tangible improvements in the health, quality of life, and survival of patients.

Rosetta Genomics, based in Rehovot, Israel and Philadelphia, Penn., is a publicly traded molecular diagnostic testing company with a focus on microRNA-based diagnostic tools designed to differentiate between various types of cancer. Through its acquisition of PersonalizeDx, the company offers core FISH, IHC (immunohistochemistry), and PCR (polymerase chain reaction) based testing capabilities and partnerships in oncology and urology that provide additional content and platforms that complement the Rosetta offerings. Rosetta’s and PersonalizeDx’s cancer testing services are commercially available through the company’s Philadelphia- and Lake Forest, Calif.-based CAP-accredited, CLIA-certified labs, respectively.

Sequenom, based in San Diego, Calif, is a publicly traded diagnostic testing and genetics analysis company focused on providing products, services, diagnostic testing, applications, and genetic analysis products that translate the results of genomic science into solutions for biomedical research, translational research, molecular medicine applications, and agricultural, livestock, and other areas of research. The company is researching, developing, and pursuing the commercialization of various noninvasive molecular diagnostic tests for prenatal genetic disorders and diseases, oncology, infectious diseases, and other diseases and disorders.

Station X, based in San Francisco, Calif., is a privately held company that develops software for scientists and clinicians who work with human genomics data in either research or clinical settings. The company’s flagship product, GenePool for Biomarker Discovery, has been designed in conjunction with a leading molecular diagnostics company. Building on GenePool technology, the company is also creating validated software applications to support clinical trials and the interpretation of comprehensive genetic test panels.

Request full report
To access the full report, please provide your contact information in the form below. Thank you for your interest in First Analysis research.
